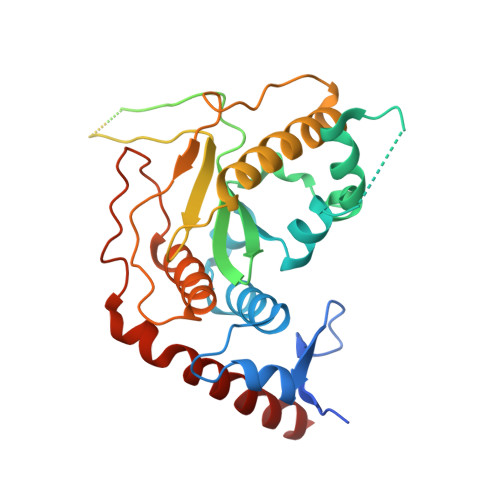
Structure of the Human SENP7 Catalytic Domain and Poly-SUMO Deconjugation Activities for SENP6 and SENP7.
Lima, C.D., Reverter, D.(2008) J Biological Chem 283: 32045-32055
- PubMed: 18799455 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M805655200
- Primary Citation Related Structures:
3EAY - PubMed Abstract:
Small ubiquitin-like modifier (SUMO) proteases regulate the abundance and lifetime of SUMO-conjugated substrates by antagonizing reactions catalyzed by SUMO-conjugating enzymes. Six SUMO proteases constitute the human SENP/ULP protease family (SENP1-3 and SENP5-7). SENP6 and SENP7 include the most divergent class of SUMO proteases, which also includes the yeast enzyme ULP2. We present the crystal structure of the SENP7 catalytic domain at a resolution of 2.4 angstroms. Comparison with structures of human SENP1 and SENP2 reveals unique elements that differ from previously characterized structures of SUMO-deconjugating enzymes. Biochemical assays show that SENP6 and SENP7 prefer SUMO2 or SUMO3 in deconjugation reactions with rates comparable with those catalyzed by SENP2, particularly during cleavage of di-SUMO2, di-SUMO3, and poly-SUMO chains composed of SUMO2 or SUMO3. In contrast, SENP6 and SENP7 exhibit lower rates for processing pre-SUMO1, pre-SUMO2, or pre-SUMO3 in comparison with SENP2. Structure-guided mutational analysis reveals elements unique to the SENP6 and SENP7 subclass of SENP/ULP proteases that contribute to protease function during deconjugation of poly-SUMO chains.
- Structural Biology Program, Sloan-Kettering Institute, New York, New York 10065, USA. limac@mskcc.org
Organizational Affiliation: