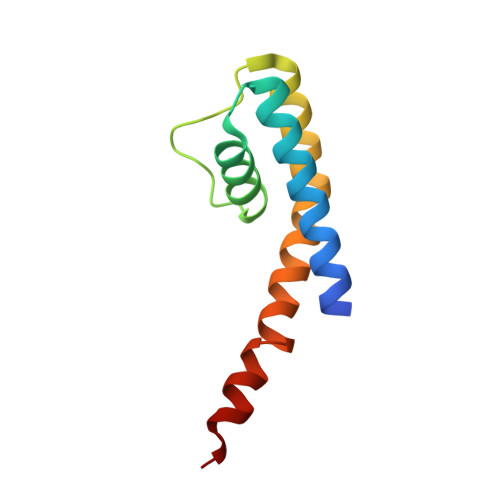
High-resolution structure of the open NaK channel
Alam, A., Jiang, Y.(2009) Nat Struct Mol Biol 16: 30-34
- PubMed: 19098917 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1531
- Primary Citation Related Structures:
3E86 - PubMed Abstract:
We report the crystal structure of the nonselective cation channel NaK from Bacillus cereus at a resolution of 1.6 A. The structure reveals the intracellular gate in an open state, as opposed to the closed form reported previously, making NaK the only channel for which the three-dimensional structures of both conformations are known. Channel opening follows a conserved mechanism of inner helix bending using a flexible glycine residue, the gating hinge, seen in MthK and most other tetrameric cation channels. Additionally, distinct inter and intrasubunit rearrangements involved in channel gating are seen and characterized for the first time along with inner helix twisting motions. Furthermore, we identify a residue deeper within the cavity of the channel pore, Phe92, which is likely to form a constriction point within the open pore, restricting ion flux through the channel. Mutating this residue to alanine causes a subsequent increase in ion-conduction rates as measured by (86)Rb flux assays. The structures of both the open and closed conformations of the NaK channel correlate well with those of equivalent K(+) channel conformations, namely MthK and KcsA, respectively.
- Department of Physiology, University of Texas Southwestern Medical Center, 5323 Harry Hines Boulevard, Dallas, Texas 75390-9040, USA.
Organizational Affiliation: