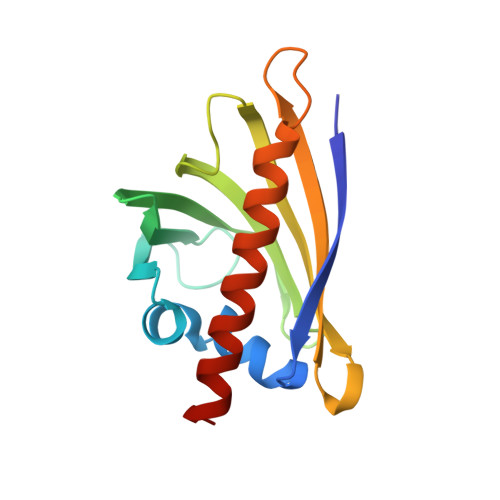
Cytokinin-induced structural adaptability of a Lupinus luteus PR-10 protein.
Fernandes, H., Bujacz, A., Bujacz, G., Jelen, F., Jasinski, M., Kachlicki, P., Otlewski, J., Sikorski, M.M., Jaskolski, M.(2009) FEBS J 276: 1596-1609
- PubMed: 19220853
- DOI: https://doi.org/10.1111/j.1742-4658.2009.06892.x
- Primary Citation Related Structures:
3E85 - PubMed Abstract:
Plant pathogenesis-related (PR) proteins of class 10 are the only group among the 17 PR protein families that are intracellular and cytosolic. Sequence conservation and the wide distribution of PR-10 proteins throughout the plant kingdom are an indication of an indispensable function in plants, but their true biological role remains obscure. Crystal and solution structures for several homologues have shown a similar overall fold with a vast internal cavity which, together with structural similarities to the steroidogenic acute regulatory protein-related lipid transfer domain and cytokinin-specific binding proteins, strongly indicate a ligand-binding role for the PR-10 proteins. This article describes the structure of a complex between a classic PR-10 protein [Lupinus luteus (yellow lupine) PR-10 protein of subclass 2, LlPR-10.2B] and N,N'-diphenylurea, a synthetic cytokinin. Synthetic cytokinins have been shown in various bioassays to exhibit activity similar to that of natural cytokinins. The present 1.95 A resolution crystallographic model reveals four N,N'-diphenylurea molecules in the hydrophobic cavity of the protein and a degree of conformational changes accompanying ligand binding. The structural adaptability of LlPR-10.2B and its ability to bind different cytokinins suggest that this protein, and perhaps other PR-10 proteins as well, can act as a reservoir of cytokinin molecules in the aqueous environment of a plant cell.
- Institute of Bioorganic Chemistry, Polish Academy of Sciences, Poznan, Poland.
Organizational Affiliation: