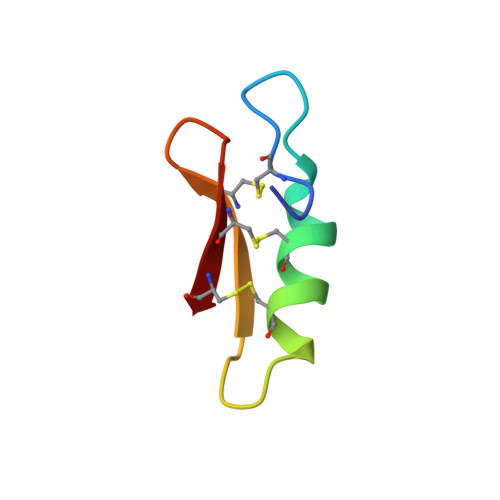
Racemic crystallography of synthetic protein enantiomers used to determine the X-ray structure of plectasin by direct methods
Mandal, K., Pentelute, B.L., Tereshko, V., Thammavongsa, V., Schneewind, O., Kossiakoff, A.A., Kent, S.B.(2009) Protein Sci 18: 1146-1154
- PubMed: 19472324 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.127
- Primary Citation Related Structures:
3E7R, 3E7U - PubMed Abstract:
We describe the use of racemic crystallography to determine the X-ray structure of the natural product plectasin, a potent antimicrobial protein recently isolated from fungus. The protein enantiomers L-plectasin and D-plectasin were prepared by total chemical synthesis; interestingly, L-plectasin showed the expected antimicrobial activity, while D-plectasin was devoid of such activity. The mirror image proteins were then used for racemic crystallization. Synchrotron X-ray diffraction data were collected to atomic resolution from a racemic plectasin crystal; the racemate crystallized in the achiral centrosymmetric space group P1 with one L-plectasin molecule and one D-plectasin molecule forming the unit cell. Dimer-like intermolecular interactions between the protein enantiomers were observed, which may account for the observed extremely low solvent content (13%-15%) and more highly ordered nature of the racemic crystals. The structure of the plectasin molecule was well defined for all 40 amino acids and was generally similar to the previously determined NMR structure, suggesting minimal impact of the crystal packing on the plectasin conformation.
- Department of Biochemistry and Molecular Biology, The University of Chicago, Chicago, IL 60637, USA. kmandal@uchicago.edu
Organizational Affiliation: