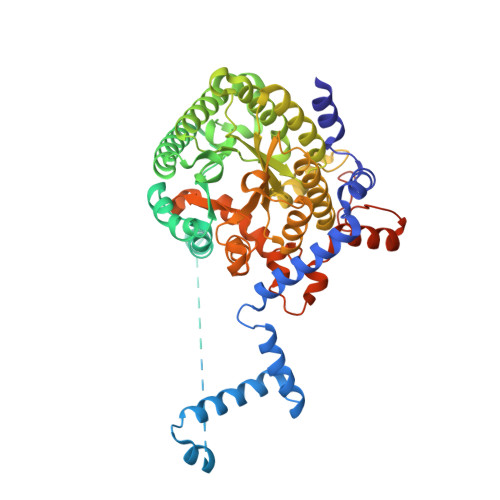
A conserved active site tyrosine residue of proline dehydrogenase helps enforce the preference for proline over hydroxyproline as the substrate.
Ostrander, E.L., Larson, J.D., Schuermann, J.P., Tanner, J.J.(2009) Biochemistry 48: 951-959
- PubMed: 19140736
- DOI: https://doi.org/10.1021/bi802094k
- Primary Citation Related Structures:
3E2Q, 3E2R, 3E2S - PubMed Abstract:
Proline dehydrogenase (PRODH) catalyzes the oxidation of l-proline to Delta-1-pyrroline-5-carboxylate. PRODHs exhibit a pronounced preference for proline over hydroxyproline (trans-4-hydroxy-l-proline) as the substrate, but the basis for specificity is unknown. The goal of this study, therefore, is to gain insight into the structural determinants of substrate specificity of this class of enzyme, with a focus on understanding how PRODHs discriminate between the two closely related molecules, proline and hydroxyproline. Two site-directed mutants of the PRODH domain of Escherichia coli PutA were created: Y540A and Y540S. Kinetics measurements were performed with both mutants. Crystal structures of Y540S complexed with hydroxyproline, proline, and the proline analogue l-tetrahydro-2-furoic acid were determined at resolutions of 1.75, 1.90, and 1.85 A, respectively. Mutation of Tyr540 increases the catalytic efficiency for hydroxyproline 3-fold and decreases the specificity for proline by factors of 20 (Y540S) and 50 (Y540A). The structures show that removal of the large phenol side chain increases the volume of the substrate-binding pocket, allowing sufficient room for the 4-hydroxyl of hydroxyproline. Furthermore, the introduced serine residue participates in recognition of hydroxyproline by forming a hydrogen bond with the 4-hydroxyl. This result has implications for understanding the substrate specificity of the related enzyme human hydroxyproline dehydrogenase, which has serine in place of tyrosine at this key active site position. The kinetic and structural results suggest that Tyr540 is an important determinant of specificity. Structurally, it serves as a negative filter for hydroxyproline by clashing with the 4-hydroxyl group of this potential substrate.
- Department of Chemistry, University of Missouri, Columbia, Missouri 65211, USA.
Organizational Affiliation: