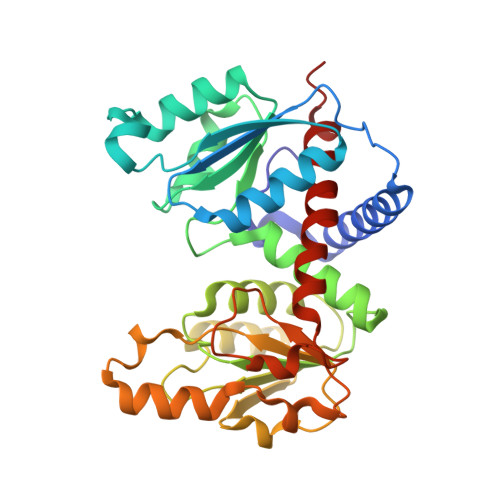
Structure of the catalytic trimer of Methanococcus jannaschii aspartate transcarbamoylase in an orthorhombic crystal form.
Vitali, J., Colaneri, M.J.(2008) Acta Crystallogr Sect F Struct Biol Cryst Commun 64: 776-780
- PubMed: 18765902 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309108025359
- Primary Citation Related Structures:
3E2P - PubMed Abstract:
Crystals of the catalytic subunit of Methanococcus jannaschii aspartate transcarbamoylase in an orthorhombic crystal form contain four crystallographically independent trimers which associate in pairs to form stable staggered complexes that are similar to each other and to a previously determined monoclinic C2 form. Each subunit has a sulfate in the central channel. The catalytic subunits in these complexes show flexibility, with the elbow angles of the monomers differing by up to 7.4 degrees between crystal forms. Moreover, there is also flexibility in the relative orientation of the trimers around their threefold axis in the complexes, with a difference of 4 degrees between crystal forms.
- Department of Physics, Cleveland State University, Euclid Avenue at East 24th Street, Cleveland, OH 44115, USA. j.vitali@csuohio.edu
Organizational Affiliation: