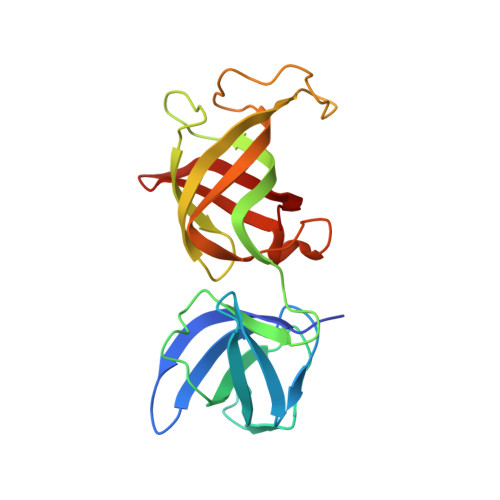
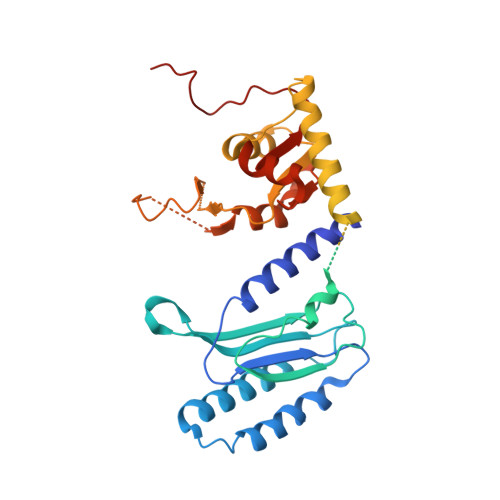
Structural insights into eRF3 and stop codon recognition by eRF1
Cheng, Z., Saito, K., Pisarev, A.V., Wada, M., Pisareva, V.P., Pestova, T.V., Gajda, M., Round, A., Kong, C., Lim, M., Nakamura, Y., Svergun, D.I., Ito, K., Song, H.(2009) Genes Dev 23: 1106-1118
- PubMed: 19417105 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/gad.1770109
- Primary Citation Related Structures:
3E1Y, 3E20 - PubMed Abstract:
Eukaryotic translation termination is mediated by two interacting release factors, eRF1 and eRF3, which act cooperatively to ensure efficient stop codon recognition and fast polypeptide release. The crystal structures of human and Schizosaccharomyces pombe full-length eRF1 in complex with eRF3 lacking the GTPase domain revealed details of the interaction between these two factors and marked conformational changes in eRF1 that occur upon binding to eRF3, leading eRF1 to resemble a tRNA molecule. Small-angle X-ray scattering analysis of the eRF1/eRF3/GTP complex suggested that eRF1's M domain contacts eRF3's GTPase domain. Consistently, mutation of Arg192, which is predicted to come in close contact with the switch regions of eRF3, revealed its important role for eRF1's stimulatory effect on eRF3's GTPase activity. An ATP molecule used as a crystallization additive was bound in eRF1's putative decoding area. Mutational analysis of the ATP-binding site shed light on the mechanism of stop codon recognition by eRF1.
- Cancer and Developmental Cell Biology Division, Institute of Molecular and Cell Biology, Agency for Science, Technology, and Research (A*STAR), Singapore, Singapore.
Organizational Affiliation: