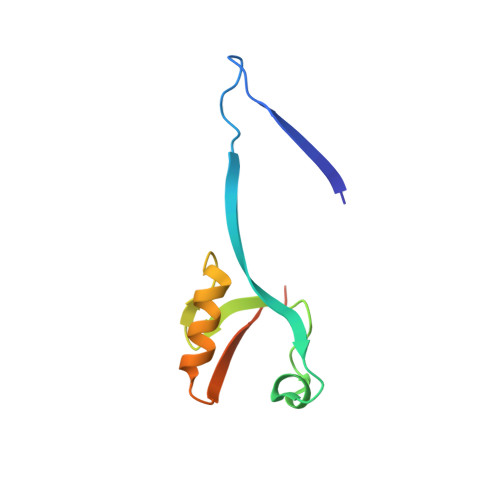
Structure of the second PDZ domain from human zonula occludens 2
Chen, H., Tong, S., Li, X., Wu, J., Zhu, Z., Niu, L., Teng, M.(2009) Acta Crystallogr Sect F Struct Biol Cryst Commun 65: 327-330
- PubMed: 19342771 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309109002334
- Primary Citation Related Structures:
3E17 - PubMed Abstract:
Human zonula occludens 2 (ZO-2) protein is a multi-domain protein that consists of an SH3 domain, a GK domain and three copies of a PDZ domain with slight divergence. The three PDZ domains act as protein-recognition modules that may mediate protein assembly and subunit localization. The crystal structure of the second PDZ domain of ZO-2 (ZO-2 PDZ2) was determined by molecular replacement at 1.75 A resolution, revealing a dimer in the asymmetric unit. The dimer is stabilized by extensive symmetrical domain-swapping of the beta1 and beta2 strands. Structural comparison shows that the ZO-2 PDZ2 homodimer may have a similar ligand-binding pattern to the ZO-1 PDZ2-connexin 43 complex.
- Hefei National Laboratory for Physical Sciences at Microscale, School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, People's Republic of China.
Organizational Affiliation: