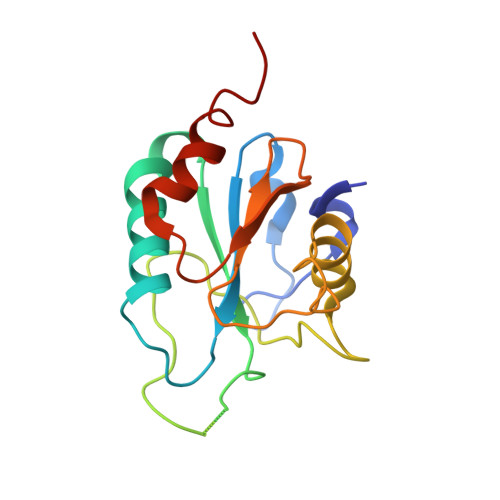
Structural insights into the catalytic mechanism of Trypanosoma cruzi GPXI (glutathione peroxidase-like enzyme I).
Patel, S., Hussain, S., Harris, R., Sardiwal, S., Kelly, J.M., Wilkinson, S.R., Driscoll, P.C., Djordjevic, S.(2010) Biochem J 425: 513-522
- PubMed: 19886864
- DOI: https://doi.org/10.1042/BJ20091167
- Primary Citation Related Structures:
3E0U - PubMed Abstract:
Current drug therapies against Trypanosoma cruzi, the causative agent of Chagas disease, have limited effectiveness and are highly toxic. T. cruzi-specific metabolic pathways that utilize trypanothione for the reduction of peroxides are being explored as potential novel therapeutic targets. In the present study we solved the X-ray crystal structure of one of the T. cruzi enzymes involved in peroxide reduction, the glutathione peroxidase-like enzyme TcGPXI (T. cruzi glutathione peroxidase-like enzyme I). We also characterized the wild-type, C48G and C96G variants of TcGPXI by NMR spectroscopy and biochemical assays. Our results show that residues Cys48 and Cys96 are required for catalytic activity. In solution, the TcGPXI molecule readily forms a Cys48-Cys96 disulfide bridge and the polypeptide segment containing Cys96 lacks regular secondary structure. NMR spectra of the reduced TcGPXI are indicative of a protein that undergoes widespread conformational exchange on an intermediate time scale. Despite the absence of the disulfide bond, the active site mutant proteins acquired an oxidized-like conformation as judged from their NMR spectra. The protein that was used for crystallization was pre-oxidized by t-butyl hydroperoxide; however, the electron density maps clearly showed that the active site cysteine residues are in the reduced thiol form, indicative of X-ray-induced reduction. Our crystallographic and solution studies suggest a level of structural plasticity in TcGPXI consistent with the requirement of the atypical two-cysteine (2-Cys) peroxiredoxin-like mechanism implied by the behaviour of the Cys48 and Cys96 mutant proteins.
- Institute of Structural and Molecular Biology, Division of Biosciences, University College London, London WC1E 6BT, UK. s.djordjevic@ucl.ac.uk
Organizational Affiliation: