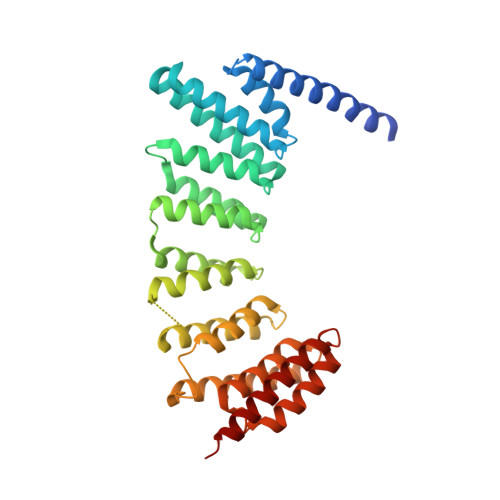
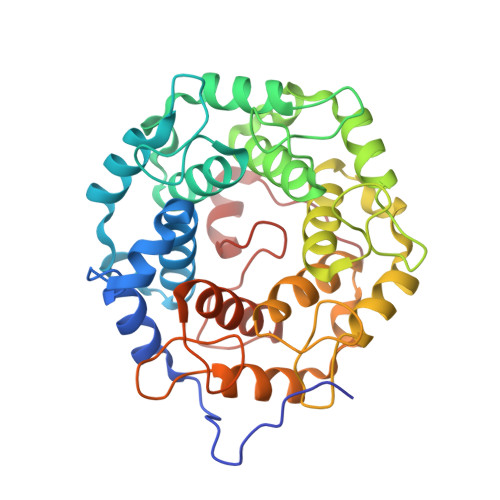
Structures of RabGGTase-substrate/product complexes provide insights into the evolution of protein prenylation
Guo, Z., Wu, Y.-W., Das, D., Delon, C., Cramer, J., Yu, S., Thuns, S., Lupilova, N., Waldmann, H., Brunsveld, L., Goody, R.S., Alexandrov, K., Blankenfeldt, W.(2008) EMBO J 27: 2444-2456
- PubMed: 18756270 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2008.164
- Primary Citation Related Structures:
3DSS, 3DST, 3DSU, 3DSV, 3DSW, 3DSX - PubMed Abstract:
Post-translational isoprenylation of proteins is carried out by three related enzymes: farnesyltransferase, geranylgeranyl transferase-I, and Rab geranylgeranyl transferase (RabGGTase). Despite the fact that the last one is responsible for the largest number of individual protein prenylation events in the cell, no structural information is available on its interaction with substrates and products. Here, we present structural and biophysical analyses of RabGGTase in complex with phosphoisoprenoids as well as with the prenylated peptides that mimic the C terminus of Rab7 GTPase. The data demonstrate that, unlike other protein prenyl transferases, both RabGGTase and its substrate RabGTPases completely 'outsource' their specificity for each other to an accessory subunit, the Rab escort protein (REP). REP mediates the placement of the C terminus of RabGTPase into the active site of RabGGTase through a series protein-protein interactions of decreasing strength and selectivity. This arrangement enables RabGGTase to prenylate any cysteine-containing sequence. On the basis of our structural and thermodynamic data, we propose that RabGGTase has evolved from a GGTase-I-like molecule that 'learned' to interact with a recycling factor (GDI) that, in turn, eventually gave rise to REP.
- Department of Physical Biochemistry, Max-Planck-Institute for Molecular Physiology, Dortmund, Germany.
Organizational Affiliation: