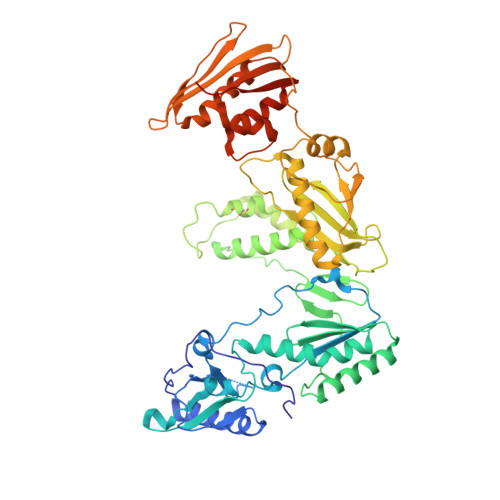
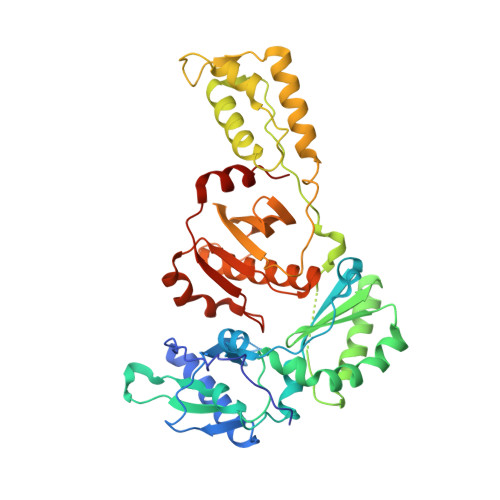
Structural basis for the improved drug resistance profile of new generation benzophenone non-nucleoside HIV-1 reverse transcriptase inhibitors.
Ren, J., Chamberlain, P.P., Stamp, A., Short, S.A., Weaver, K.L., Romines, K.R., Hazen, R., Freeman, A., Ferris, R.G., Andrews, C.W., Boone, L., Chan, J.H., Stammers, D.K.(2008) J Med Chem 51: 5000-5008
- PubMed: 18665583 Search on PubMed
- DOI: https://doi.org/10.1021/jm8004493
- Primary Citation Related Structures:
3DLE, 3DLG, 3DM2, 3DMJ, 3DOK, 3DOL - PubMed Abstract:
Owing to the emergence of resistant virus, next generation non-nucleoside HIV reverse transcriptase inhibitors (NNRTIs) with improved drug resistance profiles have been developed to treat HIV infection. Crystal structures of HIV-1 RT complexed with benzophenones optimized for inhibition of HIV mutants that were resistant to the prototype benzophenone GF128590 indicate factors contributing to the resilience of later compounds in the series (GW4511, GW678248). Meta-substituents on the benzophenone A-ring had the designed effect of inducing better contacts with the conserved W229 while reducing aromatic stacking interactions with the highly mutable Y181 side chain, which unexpectedly adopted a "down" position. Up to four main-chain hydrogen bonds to the inhibitor also appear significant in contributing to resilience. Structures of mutant RTs (K103N, V106A/Y181C) with benzophenones showed only small rearrangements of the NNRTIs relative to wild-type. Hence, adaptation to a mutated NNRTI pocket by inhibitor rearrangement appears less significant for benzophenones than other next-generation NNRTIs.
- Division of Structural Biology, The Wellcome Trust Centre for Human Genetics, Henry Wellcome Building for Genomic Medicine, University of Oxford, Roosevelt Drive, Oxford, OX3 7BN, UK.
Organizational Affiliation: