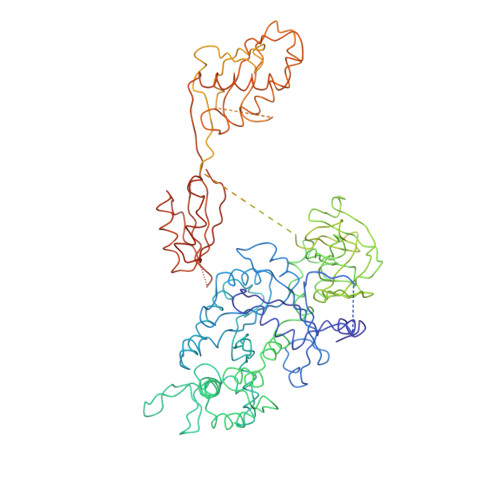
Visualization of the eEF2-80S ribosome transition-state complex by cryo-electron microscopy.
Sengupta, J., Nilsson, J., Gursky, R., Kjeldgaard, M., Nissen, P., Frank, J.(2008) J Mol Biology 382: 179-187
- PubMed: 18644383 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.07.004
- Primary Citation Related Structures:
3DNY, 3DWU - PubMed Abstract:
In an attempt to understand ribosome-induced GTP hydrolysis on eEF2, we determined a 12.6-A cryo-electron microscopy reconstruction of the eEF2-bound 80S ribosome in the presence of aluminum tetrafluoride and GDP, with aluminum tetrafluoride mimicking the gamma-phosphate during hydrolysis. This is the first visualization of a structure representing a transition-state complex on the ribosome. Tight interactions are observed between the factor's G domain and the large ribosomal subunit, as well as between domain IV and an intersubunit bridge. In contrast, some of the domains of eEF2 implicated in small subunit binding display a large degree of flexibility. Furthermore, we find support for a transition-state model conformation of the switch I region in this complex where the reoriented switch I region interacts with a conserved rRNA region of the 40S subunit formed by loops of the 18S RNA helices 8 and 14. This complex is structurally distinct from the eEF2-bound 80S ribosome complexes previously reported, and analysis of this map sheds light on the GTPase-coupled translocation mechanism.
- Wadsworth Center, New York State Department of Health, Empire State Plaza, Albany, NY 12201-0509, USA.
Organizational Affiliation: