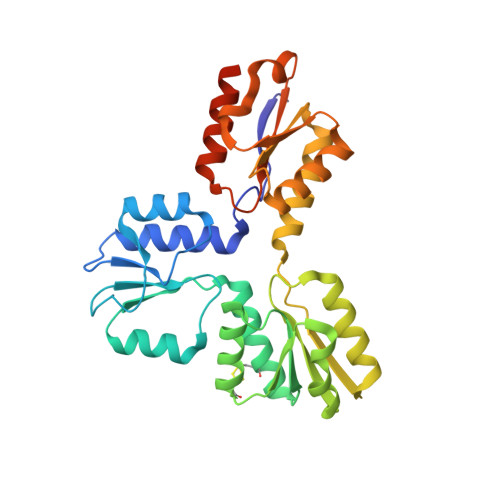
Structure of (E)-4-Hydroxy-3-methyl-but-2-enyl Diphosphate Reductase, the Terminal Enzyme of the Non-Mevalonate Pathway.
Rekittke, I., Wiesner, J., Demmer, U., Warkentin, E., Xu, W., Troschke, K., Hintz, M., No, J.H., Duin, E.C., Oldfield, E., Jomaa, H., Ermler, U.(2008) J Am Chem Soc
- PubMed: 19035630 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ja806668q
- Primary Citation Related Structures:
3DNF - PubMed Abstract:
Molecular evolution has evolved two metabolic routes for isoprenoid biosynthesis: the mevalonate and the 2-C-methyl-D-erythritol-4-phosphate (MEP) pathway. The MEP pathway is used by most pathogenic bacteria and some parasitic protozoa (including the malaria parasite, Plasmodium falciparum) as well as by plants, but is not present in animals. The terminal reaction of the MEP pathway is catalyzed by (E)-4-hydroxy-3-methyl-but-2-enyl diphosphate (HMBPP) reductase (LytB), an enzyme that converts HMBPP into isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP). Here, we present the structure of Aquifex aeolicus LytB, at 1.65 A resolution. The protein adopts a cloverleaf or trefoil-like structure with each monomer in the dimer containing three alpha/beta domains surrounding a central [Fe3S4] cluster ligated to Cys13, Cys96, and Cys193. Two highly conserved His (His 42 and His 124) and a totally conserved Glu (Glu126) are located in the same central site and are proposed to be involved in ligand binding and catalysis. Substrate access is proposed to occur from the front-side face of the protein, with the HMBPP diphosphate binding to the two His and the 4OH of HMBPP binding to the fourth iron thought to be present in activated clusters, while Glu126 provides the protons required for IPP/DMAPP formation.
- Institut für Klinische Immunologie and Transfusionsmedizin, Justus-Liebig-Universität Giessen, Langhansstrasse 7, 35385 Giessen, Germany.
Organizational Affiliation: