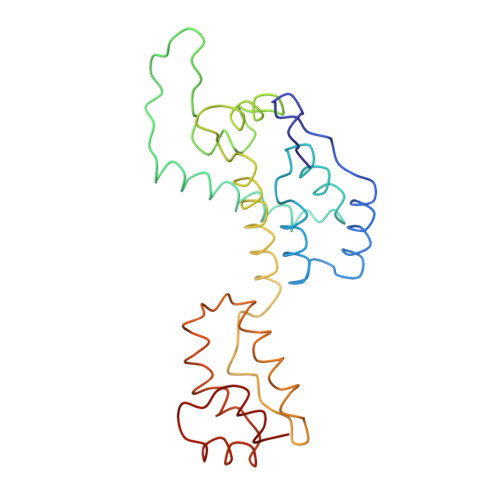
Structure of full-length HIV-1 CA: a model for the mature capsid lattice
Ganser-Pornillos, B.K., Cheng, A., Yeager, M.(2007) Cell 131: 70-79
- PubMed: 17923088 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2007.08.018
- Primary Citation Related Structures:
3DIK - PubMed Abstract:
The capsids of mature retroviruses perform the essential function of organizing the viral genome for efficient replication. These capsids are modeled as fullerene structures composed of closed hexameric arrays of the viral CA protein, but a high-resolution structure of the lattice has remained elusive. A three-dimensional map of two-dimensional crystals of the R18L mutant of HIV-1 CA was derived by electron cryocrystallography. The docking of high-resolution domain structures into the map yielded the first unambiguous model for full-length HIV-1 CA. Three important protein-protein assembly interfaces are required for capsid formation. Each CA hexamer is composed of an inner ring of six N-terminal domains and an outer ring of C-terminal domains that form dimeric linkers connecting neighboring hexamers. Interactions between the two domains of CA further stabilize the hexamer and provide a structural explanation for the mechanism of action of known HIV-1 assembly inhibitors.
- The Scripps Research Institute, Department of Cell Biology, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: