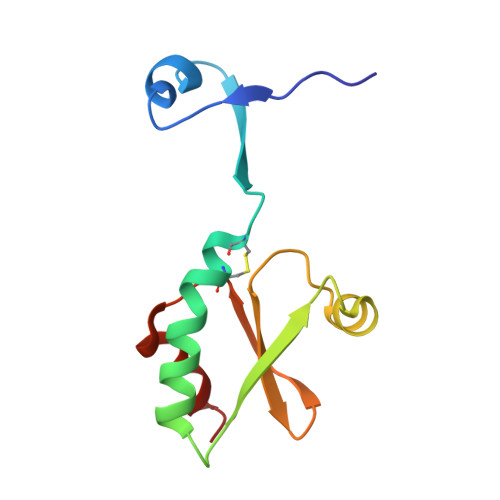
Coupling of domain swapping to kinetic stability in a thioredoxin mutant
Garcia-Pino, A., Martinez-Rodriguez, S., Wahni, K., Wyns, L., Loris, R., Messens, J.(2009) J Mol Biology 385: 1590-1599
- PubMed: 19071139 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.11.040
- Primary Citation Related Structures:
3DIE - PubMed Abstract:
The thioredoxin (Trx) fold is a small monomeric domain that is ubiquitous in redox-active enzymes. Trxs are characterized by a typical WCGPC active-site sequence motif. A single active-site mutation of the tryptophan to an alanine in Staphylococcus aureus Trx converts the oxidized protein into a biologically inactive domain-swapped dimer. While the monomeric protein unfolds reversibly in a two-state manner, the oxidized dimeric form is kinetically stable and converts to the monomeric form upon refolding. After reduction, the half-life of the dimer decreases many orders of magnitude to approximately 4.3 h, indicating that the active-site disulfide between Cys29 and Cys32 is an important determinant for the kinetics of unfolding. We propose kinetic stability as a possible evolutionary strategy in the evolution of multimeric proteins from their monomeric ancestors by domain swapping, which, for this biologically inactive Trx mutant, turned out to be an evolutionary dead end.
- Department of Molecular and Cellular Interactions, Vlaams Instituut voor Biotechnologie, Brussels, Belgium.
Organizational Affiliation: