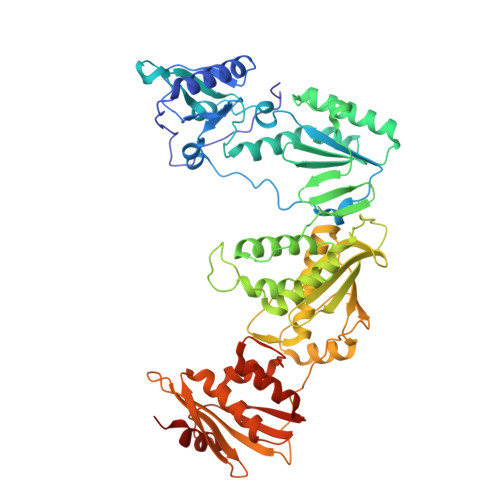
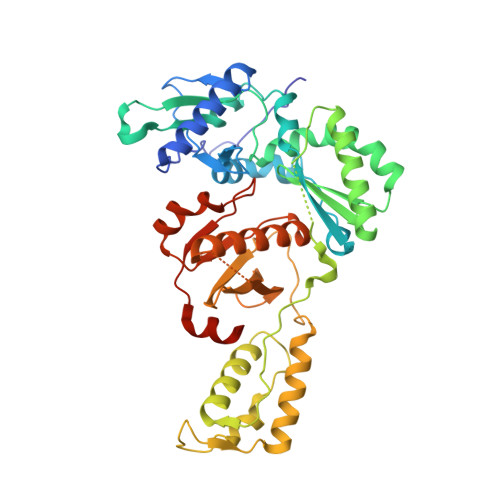
Discovery and optimization of pyridazinone non-nucleoside inhibitors of HIV-1 reverse transcriptase.
Sweeney, Z.K., Dunn, J.P., Li, Y., Heilek, G., Dunten, P., Elworthy, T.R., Han, X., Harris, S.F., Hirschfeld, D.R., Hogg, J.H., Huber, W., Kaiser, A.C., Kertesz, D.J., Kim, W., Mirzadegan, T., Roepel, M.G., Saito, Y.D., Silva, T.M., Swallow, S., Tracy, J.L., Villasenor, A., Vora, H., Zhou, A.S., Klumpp, K.(2008) Bioorg Med Chem Lett 18: 4352-4354
- PubMed: 18632268 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2008.06.072
- Primary Citation Related Structures:
3DI6 - PubMed Abstract:
A series of benzyl pyridazinones were evaluated as HIV-1 non-nucleoside reverse transcriptase inhibitors (NNRTIs). Several members of this series showed good activity against the wild-type virus and NNRTI-resistant viruses. The binding of inhibitor 5a to HIV-RT was analyzed by surface plasmon resonance spectroscopy. Pharmacokinetic studies of 5a in rat and dog demonstrated that this compound has good oral bioavailability in animal species. The crystal structure of a complex between HIV-RT and inhibitor 4c is also described.
- Roche Palo Alto LLC, 3431 Hillview Avenue, Palo Alto, CA 94304, USA. zachary.sweeney@roche.com
Organizational Affiliation: