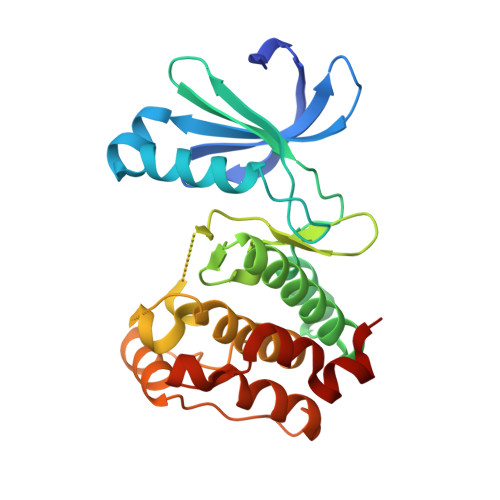
Discovery of an Aurora kinase inhibitor through site-specific dynamic combinatorial chemistry.
Cancilla, M.T., He, M.M., Viswanathan, N., Simmons, R.L., Taylor, M., Fung, A.D., Cao, K., Erlanson, D.A.(2008) Bioorg Med Chem Lett 18: 3978-3981
- PubMed: 18579375 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2008.06.011
- Primary Citation Related Structures:
3DAJ - PubMed Abstract:
We demonstrate a fragment-based lead discovery method that combines site-directed ligand discovery with dynamic combinatorial chemistry. Our technique targets dynamic combinatorial screening to a specified region of a protein by using reversible disulfide chemistry. We have used this technology to rapidly identify inhibitors of the drug target Aurora A that span the purine-binding site and the adaptive pocket of the kinase. The binding mode of a noncovalent inhibitor has been further characterized through crystallography.
- Sunesis Pharmaceuticals, Inc., 395 Oyster Point Boulevard, Suite 400, South San Francisco, CA 94080, USA.
Organizational Affiliation: