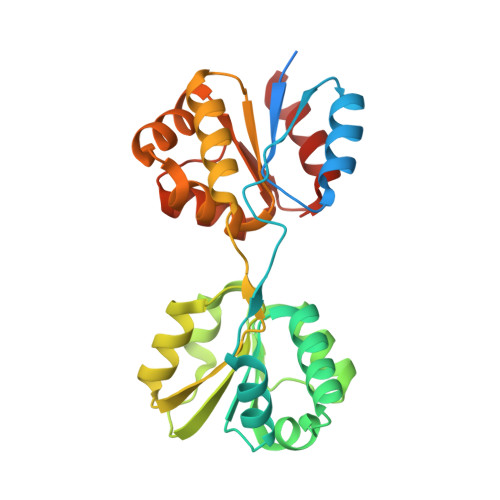
Structure and mechanistic implications of a uroporphyrinogen III synthase-product complex.
Schubert, H.L., Phillips, J.D., Heroux, A., Hill, C.P.(2008) Biochemistry 47: 8648-8655
- PubMed: 18651750 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi800635y
- Primary Citation Related Structures:
3D8N, 3D8R, 3D8S, 3D8T - PubMed Abstract:
Uroporphyrinogen III synthase (U3S) catalyzes the asymmetrical cyclization of a linear tetrapyrrole to form the physiologically relevant uroporphyrinogen III (uro'gen III) isomer during heme biosynthesis. Here, we report four apoenzyme and one product complex crystal structures of the Thermus thermophilus (HB27) U3S protein. The overlay of eight crystallographically unique U3S molecules reveals a huge range of conformational flexibility, including a "closed" product complex. The product, uro'gen III, binds between the two domains and is held in place by a network of hydrogen bonds between the product's side chain carboxylates and the protein's main chain amides. Interactions of the product A and B ring carboxylate side chains with both structural domains of U3S appear to dictate the relative orientation of the domains in the closed enzyme conformation and likely remain intact during catalysis. The product C and D rings are less constrained in the structure, consistent with the conformational changes required for the catalytic cyclization with inversion of D ring orientation. A conserved tyrosine residue is potentially positioned to facilitate loss of a hydroxyl from the substrate to initiate the catalytic reaction.
- Departments of Biochemistry and Internal Medicine, School of Medicine, University of Utah, Salt Lake City, Utah 84112, , USA. heidi@biochem.utah.edu
Organizational Affiliation: