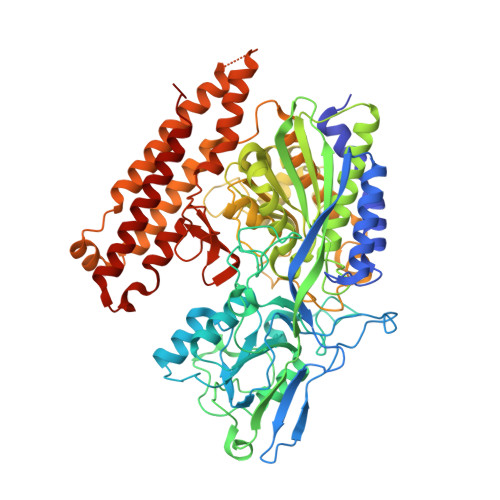
Interactions between Human Glutamate Carboxypeptidase II and Urea-Based Inhibitors: Structural Characterization
Barinka, C., Byun, Y., Dusich, C.L., Banerjee, S.R., Chen, Y., Castanares, M., Kozikowski, A.P., Mease, R.C., Pomper, M.G., Lubkowski, J.(2008) J Med Chem 51: 7737-7743
- PubMed: 19053759 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm800765e
- Primary Citation Related Structures:
3D7D, 3D7F, 3D7G, 3D7H - PubMed Abstract:
Urea-based, low molecular weight ligands of glutamate carboxypeptidase II (GCPII) have demonstrated efficacy in various models of neurological disorders and can serve as imaging agents for prostate cancer. To enhance further development of such compounds, we determined X-ray structures of four complexes between human GCPII and urea-based inhibitors at high resolution. All ligands demonstrate an invariant glutarate moiety within the S1' pocket of the enzyme. The ureido linkage between P1 and P1' inhibitor sites interacts with the active-site Zn(1)(2+) ion and the side chains of Tyr552 and His553. Interactions within the S1 pocket are defined primarily by a network of hydrogen bonds between the P1 carboxylate group of the inhibitors and the side chains of Arg534, Arg536, and Asn519. Importantly, we have identified a hydrophobic pocket accessory to the S1 site that can be exploited for structure-based design of novel GCPII inhibitors with increased lipophilicity.
- Center for Cancer Research, National Cancer Institute at Frederick, Frederick, Maryland 21702, USA.
Organizational Affiliation: