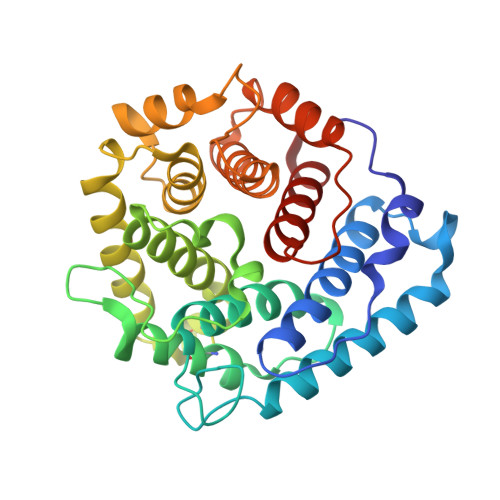
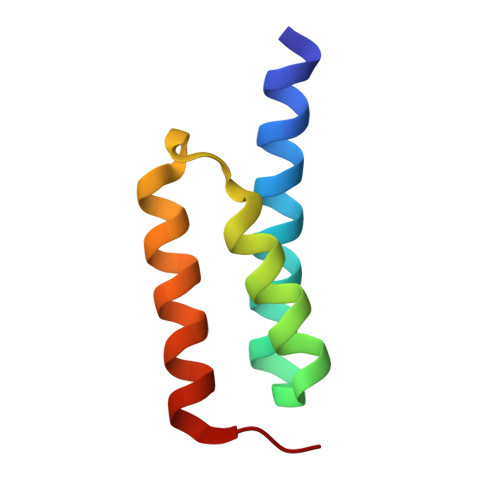
Electrostatic contributions drive the interaction between Staphylococcus aureus protein Efb-C and its complement target C3d.
Haspel, N., Ricklin, D., Geisbrecht, B.V., Kavraki, L.E., Lambris, J.D.(2008) Protein Sci 17: 1894-1906
- PubMed: 18687868 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.036624.108
- Primary Citation Related Structures:
3D5R, 3D5S - PubMed Abstract:
The C3-inhibitory domain of Staphylococcus aureus extracellular fibrinogen-binding protein (Efb-C) defines a novel three-helix bundle motif that regulates complement activation. Previous crystallographic studies of Efb-C bound to its cognate subdomain of human C3 (C3d) identified Arg-131 and Asn-138 of Efb-C as key residues for its activity. In order to characterize more completely the physical and chemical driving forces behind this important interaction, we employed in this study a combination of structural, biophysical, and computational methods to analyze the interaction of C3d with Efb-C and the single-point mutants R131A and N138A. Our results show that while these mutations do not drastically affect the structure of the Efb-C/C3d recognition complex, they have significant adverse effects on both the thermodynamic and kinetic profiles of the resulting complexes. We also characterized other key interactions along the Efb-C/C3d binding interface and found an intricate network of salt bridges and hydrogen bonds that anchor Efb-C to C3d, resulting in its potent complement inhibitory properties.
- Department of Computer Science, Rice University, Houston, Texas 77005, USA.
Organizational Affiliation: