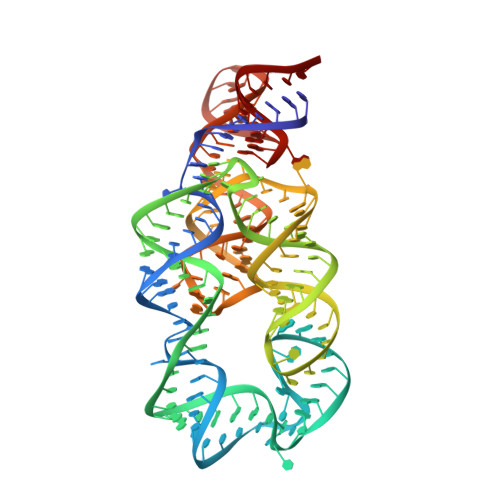
Crystal structure of the lysine riboswitch regulatory mRNA element.
Garst, A.D., Heroux, A., Rambo, R.P., Batey, R.T.(2008) J Biological Chem 283: 22347-22351
- PubMed: 18593706 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.C800120200
- Primary Citation Related Structures:
3D0U, 3D0X - PubMed Abstract:
Riboswitches are metabolite-sensitive elements found in mRNAs that control gene expression through a regulatory secondary structural switch. Along with regulation of lysine biosynthetic genes, mutations within the lysine-responsive riboswitch (L-box) play a role in the acquisition of resistance to antimicrobial lysine analogs. To understand the structural basis for lysine binding, we have determined the 2.8 angstroms resolution crystal structure of lysine bound to the Thermotoga maritima asd lysine riboswitch ligand-binding domain. The structure reveals a complex architecture scaffolding a binding pocket completely enveloping lysine. Mutations conferring antimicrobial resistance cluster around this site as well as highly conserved long range interactions, indicating that they disrupt lysine binding or proper folding of the RNA. Comparison of the free and bound forms by x-ray crystallography, small angle x-ray scattering, and chemical probing reveals almost identical structures, indicating that lysine induces only limited and local conformational changes upon binding.
- Department of Chemistry and Biochemistry, University of Colorado, Boulder, Boulder, Colorado 80309, USA.
Organizational Affiliation: