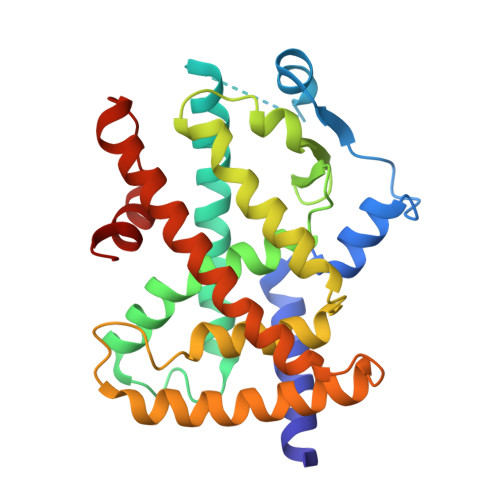
Molecular recognition of nitrated fatty acids by PPAR gamma.
Li, Y., Zhang, J., Schopfer, F.J., Martynowski, D., Garcia-Barrio, M.T., Kovach, A., Suino-Powell, K., Baker, P.R., Freeman, B.A., Chen, Y.E., Xu, H.E.(2008) Nat Struct Mol Biol 15: 865-867
- PubMed: 18604218 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1447
- Primary Citation Related Structures:
3CWD - PubMed Abstract:
Peroxisome proliferator activated receptor-gamma (PPAR gamma) regulates metabolic homeostasis and adipocyte differentiation, and it is activated by oxidized and nitrated fatty acids. Here we report the crystal structure of the PPAR gamma ligand binding domain bound to nitrated linoleic acid, a potent endogenous ligand of PPAR gamma. Structural and functional studies of receptor-ligand interactions reveal the molecular basis of PPAR gamma discrimination of various naturally occurring fatty acid derivatives.
- Department of Pharmaceutical Sciences, Center for Pharmacogenetics, 709 Salk Hall, University of Pittsburgh, Pittsburgh, Pennsylvania 15261, USA. yol21@pitt.edu
Organizational Affiliation: