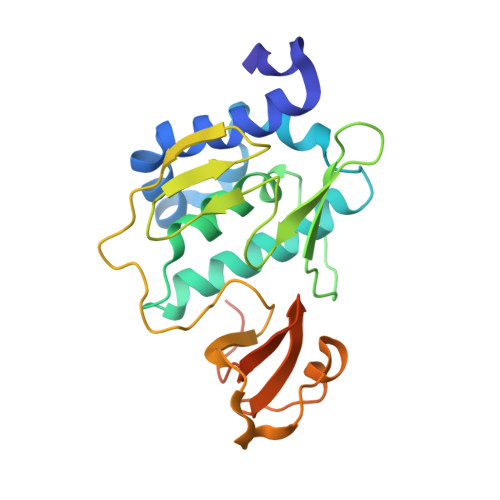
Structural basis of the influenza A virus RNA polymerase PB2 RNA-binding domain containing the pathogenicity-determinant lysine 627 residue
Kuzuhara, T., Kise, D., Yoshida, H., Horita, T., Murazaki, Y., Nishimura, A., Echigo, N., Utsunomiya, H., Tsuge, H.(2009) J Biological Chem 284: 6855-6860
- PubMed: 19144639 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.C800224200
- Primary Citation Related Structures:
3CW4 - PubMed Abstract:
Because the influenza A virus has an RNA genome, its RNA-dependent RNA polymerase, comprising the PA, PB1, and PB2 subunits, is essential for viral transcription and replication. The binding of RNA primers/promoters to the polymerases is an initiation step in viral transcription. In our current study, we reveal the 2.7 A tertiary structure of the C-terminal RNA-binding domain of PB2 by x-ray crystallography. This domain incorporates lysine 627 of PB2, and this residue is associated with the high pathogenicity and host range restriction of influenza A virus. We found from our current analyses that this lysine is located in a unique "phi"-shaped structure consisting of a helix and an encircled loop within the PB2 domain. By electrostatic analysis, we identified a highly basic groove along with this phi loop and found that lysine 627 is located in the phi loop. A PB2 domain mutant in which glutamic acid is substituted at position 627 shows significantly lower RNA binding activity. This is the first report to show a relationship between RNA binding activity and the pathogenicity-determinant lysine 627. Using the Matras program for protein three-dimensional structural comparisons, we further found that the helix bundles in the PB2 domain are similar to that of activator 1, the 40-kDa subunit of DNA replication clamp loader (replication factor C), which is also an RNA-binding protein. This suggests a functional and structural relationship between the RNA-binding mechanisms underlying both influenza A viral transcription and cellular DNA replication. Our present results thus provide important new information for developing novel drugs that target the primer/promoter RNA binding of viral RNA polymerases.
- Laboratory of Biochemistry, Faculty of Pharmaceutical Sciences, and Institute for Health Sciences, Tokushima Bunri University, Tokushima 770-8514, Japan. kuzuhara@ph.bunri-u.ac.jp
Organizational Affiliation: