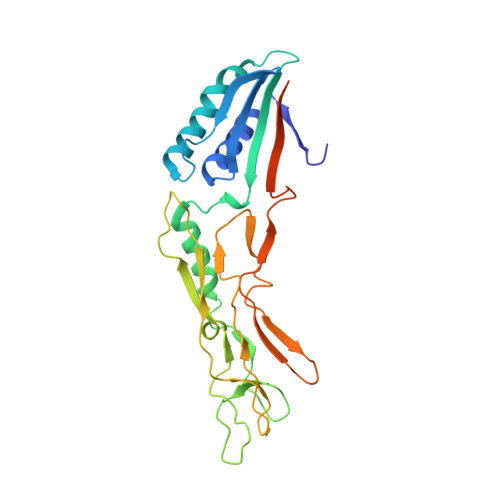
Structural insights into the molecular organization of the S-layer from Clostridium difficile
Fagan, R.P., Albesa-Jove, D., Qazi, O., Svergun, D.I., Brown, K.A., Fairweather, N.F.(2009) Mol Microbiol 71: 1308-1322
- PubMed: 19183279 Search on PubMed
- DOI: https://doi.org/10.1111/j.1365-2958.2009.06603.x
- Primary Citation Related Structures:
3CVZ - PubMed Abstract:
Clostridium difficile expresses a surface layer (S-layer) which coats the surface of the bacterium and acts as an adhesin facilitating interaction of the bacterium with host enteric cells. The S-layer contains a high-molecular-weight S-layer protein (HMW SLP) and its low-molecular-weight partner protein (LMW SLP). We show that these proteins form a tightly associated non-covalent complex, the H/L complex, and we identify the regions of both proteins responsible for complex formation. The 2.4 A X-ray crystal structure of a truncated derivative of the LMW SLP reveals two domains. Domain 1 has a two-layer sandwich architecture while domain 2, predicted to orientate towards the external environment, contains a novel fold. Small-angle X-ray scattering analysis of the H/L complex shows an elongated molecule, with the two SLPs arranged 'end-to-end' interacting with each other through a small contact area. Alignment of LMW SLPs, which exhibit high sequence diversity, reveals a core of conserved residues that could reflect functional conservation, while allowing for immune evasion through sequence variation. These structures are the first described for the S-layer of a bacterial pathogen, and provide insights into the assembly and biogenesis of the S-layer.
- Division of Cell and Molecular Biology, Centre for Molecular Microbiology and Infection, Imperial College London, London SW72AZ, UK.
Organizational Affiliation: