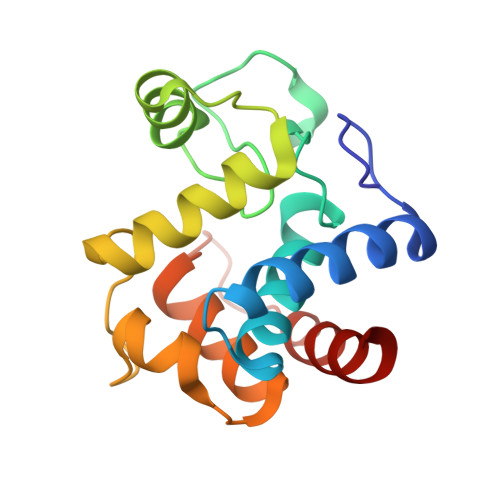
Crystal and cryoEM structural studies of a cell wall degrading enzyme in the bacteriophage phi29 tail.
Xiang, Y., Morais, M.C., Cohen, D.N., Bowman, V.D., Anderson, D.L., Rossmann, M.G.(2008) Proc Natl Acad Sci U S A 105: 9552-9557
- PubMed: 18606992 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0803787105
- Primary Citation Related Structures:
3CSQ, 3CSR, 3CSZ, 3CT0, 3CT1, 3CT5 - PubMed Abstract:
The small bacteriophage phi29 must penetrate the approximately 250-A thick external peptidoglycan cell wall and cell membrane of the Gram-positive Bacillus subtilis, before ejecting its dsDNA genome through its tail into the bacterial cytoplasm. The tail of bacteriophage phi29 is noncontractile and approximately 380 A long. A 1.8-A resolution crystal structure of gene product 13 (gp13) shows that this tail protein has spatially well separated N- and C-terminal domains, whose structures resemble lysozyme-like enzymes and metallo-endopeptidases, respectively. CryoEM reconstructions of the WT bacteriophage and mutant bacteriophages missing some or most of gp13 shows that this enzyme is located at the distal end of the phi29 tail knob. This finding suggests that gp13 functions as a tail-associated, peptidoglycan-degrading enzyme able to cleave both the polysaccharide backbone and peptide cross-links of the peptidoglycan cell wall. Comparisons of the gp13(-) mutants with the phi29 mature and emptied phage structures suggest the sequence of events that occur during the penetration of the tail through the peptidoglycan layer.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907, USA.
Organizational Affiliation: