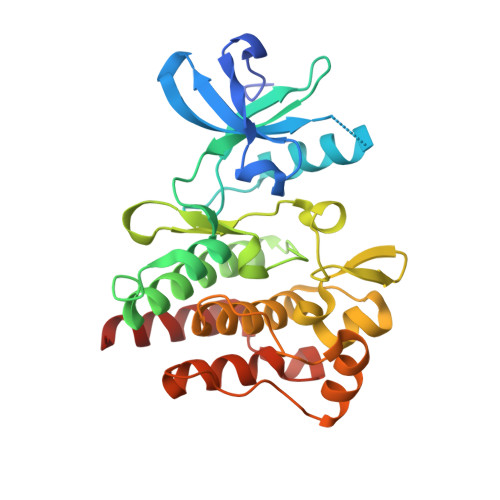
Characterization of AMN107, a selective inhibitor of native and mutant Bcr-Abl
Weisberg, E., Manley, P.W., Breitenstein, W., Brueggen, J., Cowan-Jacob, S.W., Ray, A., Huntly, B., Fabbro, D., Fendrich, G., Hall-Meyers, E., Kung, A.L., Mestan, J., Daley, G.Q., Callahan, L., Catley, L., Cavazza, C., Azam, M., Neuberg, D., Wright, R.D., Gilliland, D.G., Griffin, J.D.(2005) Cancer Cell 7: 129-141
- PubMed: 15710326 Search on PubMed
- DOI: https://doi.org/10.1016/j.ccr.2005.01.007
- Primary Citation Related Structures:
3CS9 - PubMed Abstract:
The Bcr-Abl tyrosine kinase oncogene causes chronic myelogenous leukemia (CML) and Philadelphia chromosome-positive (Ph+) acute lymphoblastic leukemia (ALL). We describe a novel selective inhibitor of Bcr-Abl, AMN107 (IC50 <30 nM), which is significantly more potent than imatinib, and active against a number of imatinib-resistant Bcr-Abl mutants. Crystallographic analysis of Abl-AMN107 complexes provides a structural explanation for the differential activity of AMN107 and imatinib against imatinib-resistant Bcr-Abl. Consistent with its in vitro and pharmacokinetic profile, AMN107 prolonged survival of mice injected with Bcr-Abl-transformed hematopoietic cell lines or primary marrow cells, and prolonged survival in imatinib-resistant CML mouse models. AMN107 is a promising new inhibitor for the therapy of CML and Ph+ ALL.
- Dana-Farber Cancer Institute, Boston, Massachusetts 02115, USA.
Organizational Affiliation: