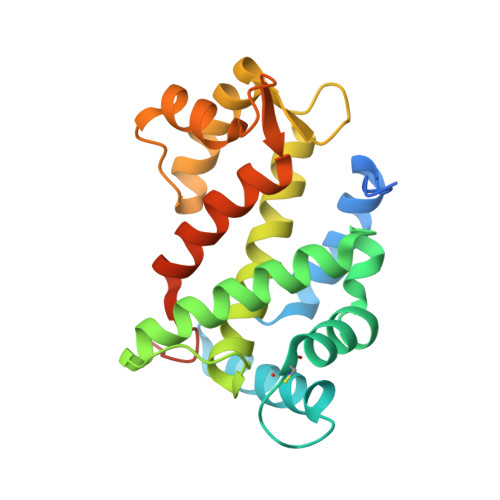
Structural Insights into Membrane Targeting by the Flagellar Calcium-binding Protein (FCaBP), a Myristoylated and Palmitoylated Calcium Sensor in Trypanosoma cruzi.
Wingard, J.N., Ladner, J., Vanarotti, M., Fisher, A.J., Robinson, H., Buchanan, K.T., Engman, D.M., Ames, J.B.(2008) J Biological Chem 283: 23388-23396
- PubMed: 18559337 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M803178200
- Primary Citation Related Structures:
3CS1 - PubMed Abstract:
The flagellar calcium-binding protein (FCaBP) of the protozoan Trypanosoma cruzi is targeted to the flagellar membrane where it regulates flagellar function and assembly. As a first step toward understanding the Ca(2+)-induced conformational changes important for membrane-targeting, we report here the x-ray crystal structure of FCaBP in the Ca(2+)-free state determined at 2.2A resolution. The first 17 residues from the N terminus appear unstructured and solvent-exposed. Residues implicated in membrane targeting (Lys-19, Lys-22, and Lys-25) are flanked by an exposed N-terminal helix (residues 26-37), forming a patch of positive charge on the protein surface that may interact electrostatically with flagellar membrane targets. The four EF-hands in FCaBP each adopt a "closed conformation" similar to that seen in Ca(2+)-free calmodulin. The overall fold of FCaBP is closest to that of grancalcin and other members of the penta EF-hand superfamily. Unlike the dimeric penta EF-hand proteins, FCaBP lacks a fifth EF-hand and is monomeric. The unstructured N-terminal region of FCaBP suggests that its covalently attached myristoyl group at the N terminus may be solvent-exposed, in contrast to the highly sequestered myristoyl group seen in recoverin and GCAP1. NMR analysis demonstrates that the myristoyl group attached to FCaBP is indeed solvent-exposed in both the Ca(2+)-free and Ca(2+)-bound states, and myristoylation has no effect on protein structure and folding stability. We propose that exposed acyl groups at the N terminus may anchor FCaBP to the flagellar membrane and that Ca(2+)-induced conformational changes may control its binding to membrane-bound protein targets.
- Center for Advanced Research in Biotechnology, University of Maryland, National Institute of Standards and Technology, Rockville, Maryland 20850, USA.
Organizational Affiliation: