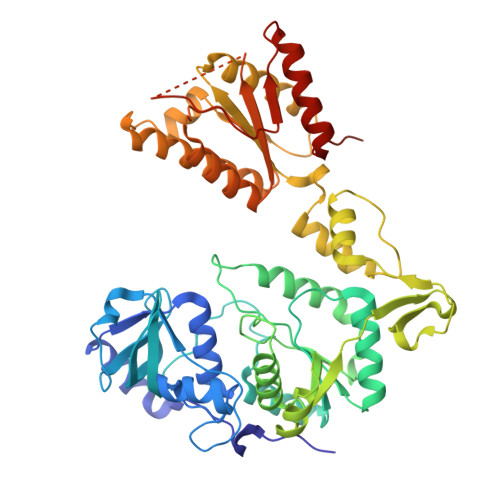
Structure of the two-domain hexameric APS kinase from Thiobacillus denitrificans: structural basis for the absence of ATP sulfurylase activity.
Gay, S.C., Segel, I.H., Fisher, A.J.(2009) Acta Crystallogr D Biol Crystallogr 65: 1021-1031
- PubMed: 19770499 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444909026547
- Primary Citation Related Structures:
3CR8 - PubMed Abstract:
The Tbd_0210 gene of the chemolithotrophic bacterium Thiobacillus denitrificans is annotated to encode a 60.5 kDa bifunctional enzyme with ATP sulfurylase and APS kinase activity. This putative bifunctional enzyme was cloned, expressed and structurally characterized. The 2.95 A resolution X-ray crystal structure reported here revealed a hexameric assembly with D(3) symmetry. Each subunit contains a large N-terminal sulfurylase-like domain and a C-terminal APS kinase domain reminiscent of the two-domain fungal ATP sulfurylases of Penicillium chrysogenum and Saccharomyces cerevisiae, which also exhibit a hexameric assembly. However, the T. denitrificans enzyme exhibits numerous structural and sequence differences in the N-terminal domain that render it inactive with respect to ATP sulfurylase activity. Surprisingly, the C-terminal domain does indeed display APS kinase activity, indicating that this gene product is a true APS kinase. Therefore, these results provide the first structural insights into a unique hexameric APS kinase that contains a nonfunctional ATP sulfurylase-like domain of unknown function.
- Department of Chemistry, University of California, One Shields Avenue, Davis, CA 95616, USA.
Organizational Affiliation: