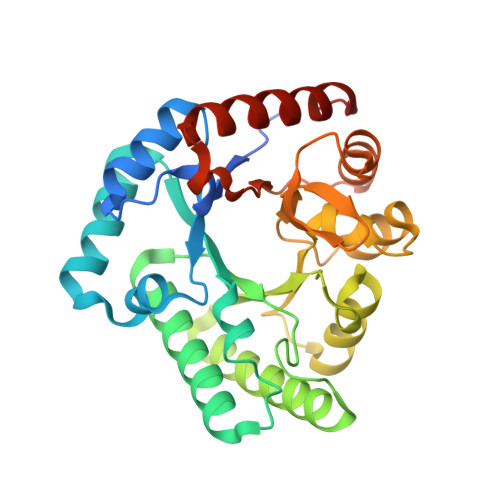
Structure of L-xylulose-5-Phosphate 3-epimerase (UlaE) from the anaerobic L-ascorbate utilization pathway of Escherichia coli: identification of a novel phosphate binding motif within a TIM barrel fold.
Shi, R., Pineda, M., Ajamian, E., Cui, Q., Matte, A., Cygler, M.(2008) J Bacteriol 190: 8137-8144
- PubMed: 18849419 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.01049-08
- Primary Citation Related Structures:
3CQH, 3CQI, 3CQJ, 3CQK - PubMed Abstract:
Three catabolic enzymes, UlaD, UlaE, and UlaF, are involved in a pathway leading to fermentation of l-ascorbate under anaerobic conditions. UlaD catalyzes a beta-keto acid decarboxylation reaction to produce L-xylulose-5-phosphate, which undergoes successive epimerization reactions with UlaE (L-xylulose-5-phosphate 3-epimerase) and UlaF (L-ribulose-5-phosphate 4-epimerase), yielding D-xylulose-5-phosphate, an intermediate in the pentose phosphate pathway. We describe here crystallographic studies of UlaE from Escherichia coli O157:H7 that complete the structural characterization of this pathway. UlaE has a triosephosphate isomerase (TIM) barrel fold and forms dimers. The active site is located at the C-terminal ends of the parallel beta-strands. The enzyme binds Zn(2+), which is coordinated by Glu155, Asp185, His211, and Glu251. We identified a phosphate-binding site formed by residues from the beta1/alpha1 loop and alpha3' helix in the N-terminal region. This site differs from the well-characterized phosphate-binding motif found in several TIM barrel superfamilies that is located at strands beta7 and beta8. The intrinsic flexibility of the active site region is reflected by two different conformations of loops forming part of the substrate-binding site. Based on computational docking of the L-xylulose 5-phosphate substrate to UlaE and structural similarities of the active site of this enzyme to the active sites of other epimerases, a metal-dependent epimerization mechanism for UlaE is proposed, and Glu155 and Glu251 are implicated as catalytic residues. Mutation and activity measurements for structurally equivalent residues in related epimerases supported this mechanistic proposal.
- Department of Biochemistry, McGill University, Montréal, Québec, Canada.
Organizational Affiliation: