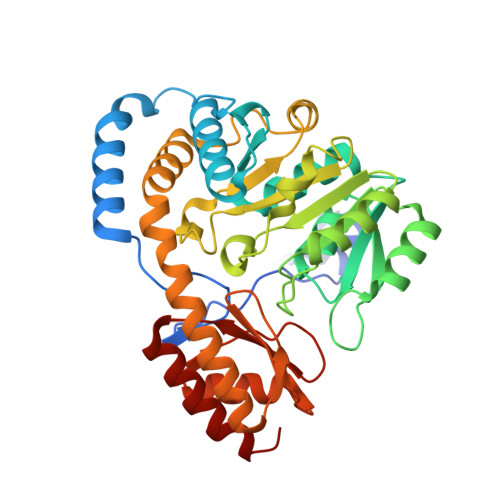
Insights into the structural basis of substrate recognition by histidinol-phosphate aminotransferase from Corynebacterium glutamicum
Marienhagen, J., Sandalova, T., Sahm, H., Eggeling, L., Schneider, G.(2008) Acta Crystallogr D Biol Crystallogr 64: 675-685
- PubMed: 18560156 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444908009438
- Primary Citation Related Structures:
3CQ4, 3CQ5, 3CQ6 - PubMed Abstract:
Histidinol-phosphate aminotransferase (HisC) is a pyridoxal 5'-phosphate-dependent enzyme that catalyzes the reversible transamination reaction between histidinol phosphate (His-P) and 2-oxoglutarate (O-Glu). The crystal structures of apo histidinol-phosphate aminotransferase from Corynebacterium glutamicum, of the internal PLP aldimine adduct and of a pyridoxamine 5-phosphate-enzyme complex were determined at resolutions of 2.2, 2.1 and 1.8 A, respectively. Residues important for substrate specificity were identified by modelling His-P into the active site and comparison with crystal structures of HisC from Thermotoga maritima and Escherichia coli. Four of the residues lining the substrate-binding pocket were studied by site-directed mutagenesis. Kinetic analysis of the Tyr21Phe mutant suggested that the hydrogen bond between the side chain of this residue and the phosphate group of His-P is important for recognition of the natural substrate and discrimination against other potential amino donors such as phenylalanine and leucine. The mutagenesis studies further indicated that residue Asn99 does not contribute to the specific recognition of the amino-acid donor, but may be involved in binding of the phosphate group of pyridoxal 5'-phosphate. The conserved residues Tyr123 and Tyr257 interact with the substrate through van der Waals interactions and their potential for hydrogen-bonding interactions is not utilized in substrate recognition, as the corresponding phenylalanine mutants show only a moderate effect on the catalytic efficiency kcat/Km.
- Institute for Biotechnology, Research Center Juelich, 52425 Juelich, Germany.
Organizational Affiliation: