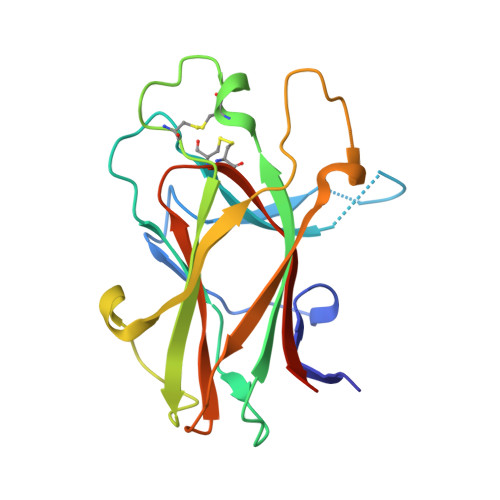
Crystal Structure and NMR Binding Reveal That Two Small Molecule Antagonists Target the High Affinity Ephrin-binding Channel of the EphA4 Receptor.
Qin, H., Shi, J., Noberini, R., Pasquale, E.B., Song, J.(2008) J Biological Chem 283: 29473-29484
- PubMed: 18708347 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M804114200
- Primary Citation Related Structures:
3CKH - PubMed Abstract:
The Eph receptor tyrosine kinases regulate a variety of physiological and pathological processes not only during development but also in adult organs, and therefore they represent a promising class of drug targets. The EphA4 receptor plays important roles in the inhibition of the regeneration of injured axons, synaptic plasticity, platelet aggregation, and likely in certain types of cancer. Here we report the first crystal structure of the EphA4 ligand-binding domain, which adopts the same jellyroll beta-sandwich architecture as shown previously for EphB2 and EphB4. The similarity with EphB receptors is high in the core beta-stranded regions, whereas large variations exist in the loops, particularly the D-E and J-K loops, which form the high affinity ephrin binding channel. We also used isothermal titration calorimetry, NMR spectroscopy, and computational docking to characterize the binding to EphA4 of two small molecules, 4- and 5-(2,5 dimethyl-pyrrol-1-yl)-2-hydroxybenzoic acid which antagonize ephrin-induced effects in EphA4-expressing cells. We show that the two molecules bind to the EphA4 ligand-binding domain with K(d) values of 20.4 and 26.4 microm, respectively. NMR heteronuclear single quantum coherence titrations revealed that upon binding, both molecules significantly perturb EphA4 residues Ile(31)-Met(32) in the D-E loop, Gln(43) in the E beta-strand, and Ile(131)-Gly(132) in the J-K loop. Molecular docking shows that they can occupy a cavity in the high affinity ephrin binding channel of EphA4 in a similar manner, by interacting mainly with the EphA4 residues in the E strand and D-E and J-K loops. However, many of the interactions observed in Eph receptor-ephrin complexes are absent, which is consistent with the small size of the two molecules and may account for their relatively weak binding affinity. Thus, our studies provide the first published structure of the ligand-binding domain of an EphA receptor of the A subclass. Furthermore, the results demonstrate that the high affinity ephrin binding channel of the Eph receptors is amenable to targeting with small molecule antagonists and suggest avenues for further optimization.
- Department of Biological Sciences, Faculty of Science, National University of Singapore, Singapore 11926.
Organizational Affiliation: