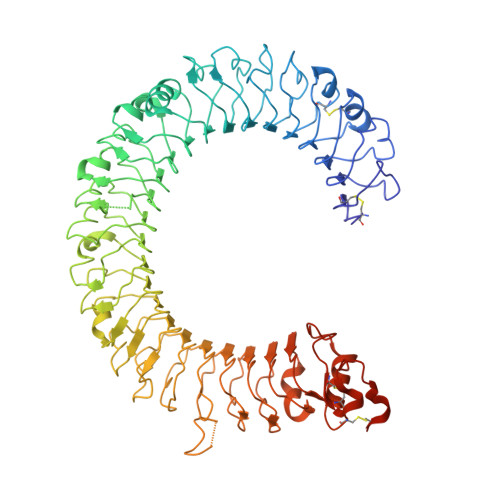
Structural basis of toll-like receptor 3 signaling with double-stranded RNA.
Liu, L., Botos, I., Wang, Y., Leonard, J.N., Shiloach, J., Segal, D.M., Davies, D.R.(2008) Science 320: 379-381
- PubMed: 18420935 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1155406
- Primary Citation Related Structures:
3CIG, 3CIY - PubMed Abstract:
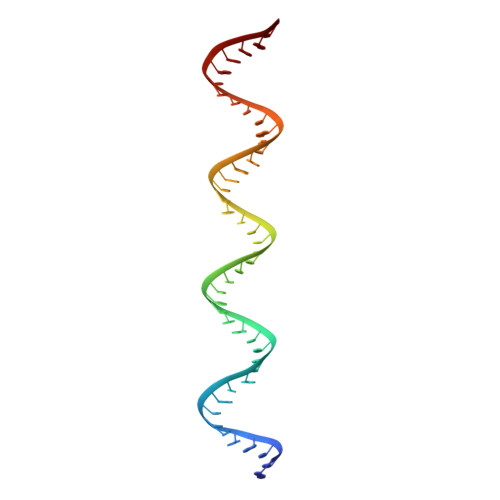
Toll-like receptor 3 (TLR3) recognizes double-stranded RNA (dsRNA), a molecular signature of most viruses, and triggers inflammatory responses that prevent viral spread. TLR3 ectodomains (ECDs) dimerize on oligonucleotides of at least 40 to 50 base pairs in length, the minimal length required for signal transduction. To establish the molecular basis for ligand binding and signaling, we determined the crystal structure of a complex between two mouse TLR3-ECDs and dsRNA at 3.4 angstrom resolution. Each TLR3-ECD binds dsRNA at two sites located at opposite ends of the TLR3 horseshoe, and an intermolecular contact between the two TLR3-ECD C-terminal domains coordinates and stabilizes the dimer. This juxtaposition could mediate downstream signaling by dimerizing the cytoplasmic Toll interleukin-1 receptor (TIR) domains. The overall shape of the TLR3-ECD does not change upon binding to dsRNA.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: