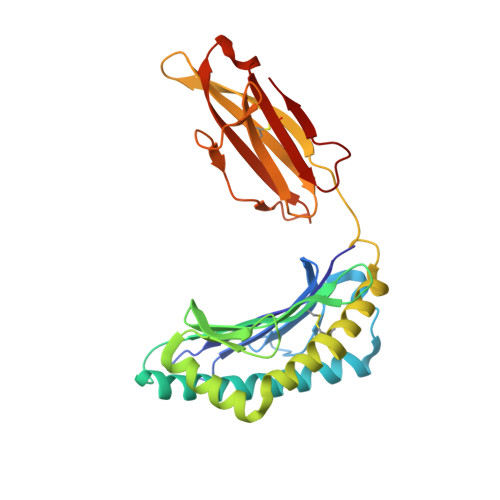
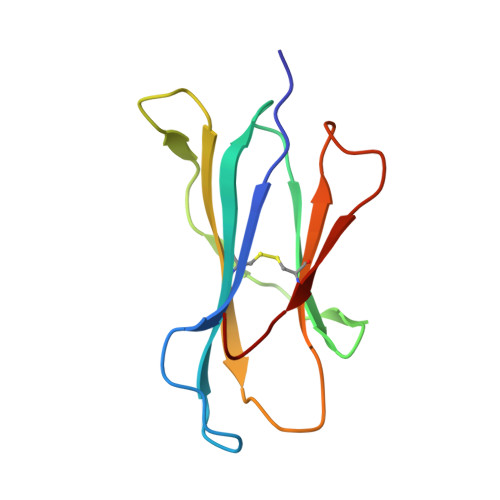
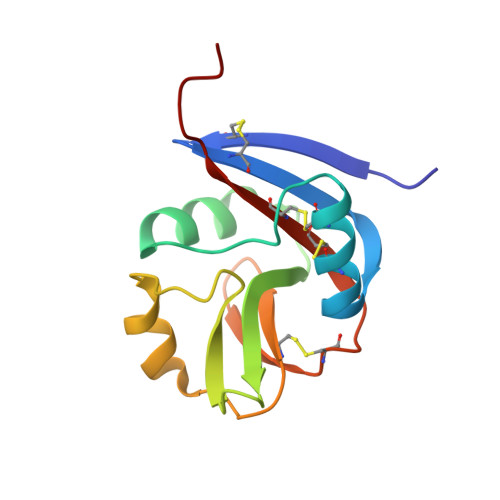
Molecular architecture of the major histocompatibility complex class I-binding site of Ly49 natural killer cell receptors.
Deng, L., Cho, S., Malchiodi, E.L., Kerzic, M.C., Dam, J., Mariuzza, R.A.(2008) J Biological Chem 283: 16840-16849
- PubMed: 18426793 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M801526200
- Primary Citation Related Structures:
3C8J, 3C8K, 3CAD - PubMed Abstract:
Natural killer (NK) cells play a vital role in the detection and destruction of virally infected and tumor cells during innate immune responses. The highly polymorphic Ly49 family of NK receptors regulates NK cell function by sensing major histocompatibility complex class I (MHC-I) molecules on target cells. Despite the determination of two Ly49-MHC-I complex structures, the molecular features of Ly49 receptors that confer specificity for particular MHC-I alleles have not been identified. To understand the functional architecture of Ly49-binding sites, we determined the crystal structures of Ly49C and Ly49G and completed refinement of the Ly49C-H-2K(b) complex. This information, combined with mutational analysis of Ly49A, permitted a structure-based classification of Ly49s that we used to dissect the binding site into three distinct regions, each having different roles in MHC recognition. One region, located at the center of the binding site, has a similar structure across the Ly49 family and mediates conserved interactions with MHC-I that contribute most to binding. However, the preference of individual Ly49s for particular MHC-I molecules is governed by two regions that flank the central region and are structurally more variable. One of the flanking regions divides Ly49s into those that recognize both H-2D and H-2K versus only H-2D ligands, whereas the other discriminates among H-2D or H-2K alleles. The modular design of Ly49-binding sites provides a framework for predicting the MHC-binding specificity of Ly49s that have not been characterized experimentally.
- Center for Advanced Research in Biotechnology, W. M. Keck Laboratory for Structural Biology, University of Maryland Biotechnology Institute, Rockville, Maryland 20850, USA.
Organizational Affiliation: