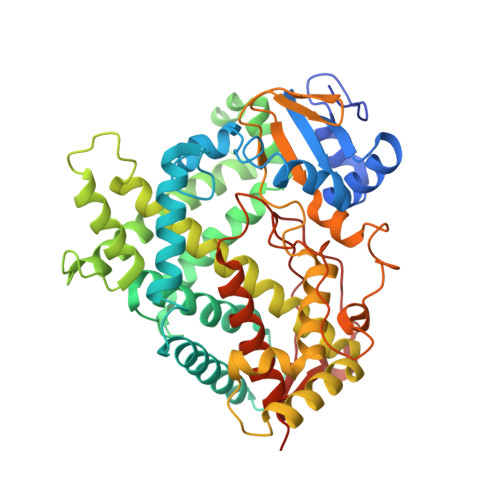
Structural analysis of CYP2R1 in complex with vitamin D3.
Strushkevich, N., Usanov, S.A., Plotnikov, A.N., Jones, G., Park, H.W.(2008) J Mol Biology 380: 95-106
- PubMed: 18511070 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.03.065
- Primary Citation Related Structures:
3C6G - PubMed Abstract:
The activation of vitamin D to its hormonal form is mediated by cytochrome P450 enzymes. CYP2R1 catalyzes the initial step converting vitamin D into 25-hydroxyvitamin D. A CYP2R1 gene mutation causes an inherited form of rickets due to 25-hydroxylase deficiency. To understand the narrow substrate specificity of CYP2R1 we obtained the hemeprotein in a highly purified state, confirmed the enzyme as a vitamin D 25-hydroxylase, and solved the crystal structure of CYP2R1 in complex with vitamin D3. The CYP2R1 structure adopts a closed conformation with the substrate access channel being covered by the ordered B'-helix and slightly opened to the surface, which defines the substrate entrance point. The active site is lined by conserved, mostly hydrophobic residues. Vitamin D3 is bound in an elongated conformation with the aliphatic side-chain pointing toward the heme. The structure reveals the secosteroid binding mode in an extended active site and allows rationalization of the molecular basis of the inherited rickets associated with CYP2R1.
- Structural Genomics Consortium, University of Toronto, Toronto, Ontario, Canada.
Organizational Affiliation: