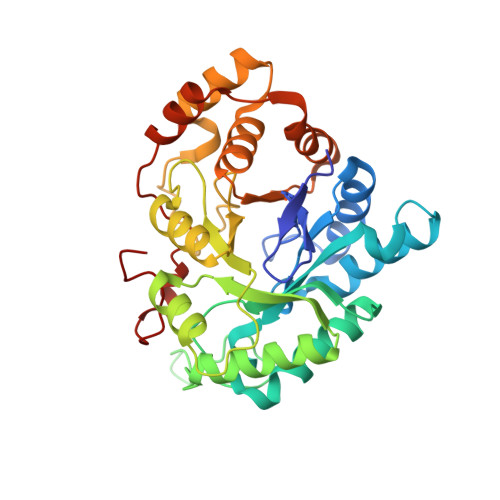
Selectivity determinants of inhibitor binding to human 20alpha-hydroxysteroid dehydrogenase: crystal structure of the enzyme in ternary complex with coenzyme and the potent inhibitor 3,5-dichlorosalicylic acid
Dhagat, U., Endo, S., Sumii, R., Hara, A., El-Kabbani, O.(2008) J Med Chem 51: 4844-4848
- PubMed: 18620380 Search on PubMed
- DOI: https://doi.org/10.1021/jm8003575
- Primary Citation Related Structures:
3C3U - PubMed Abstract:
The crystal structure of human 20alpha-hydroxysteroid dehydrogenase (AKR1C1) in ternary complex with the coenzyme NADP (+) and the potent inhibitor 3,5-dichlorosalicylic acid was determined at a resolution of 1.8 A. The inhibitor is held in place by a network of hydrogen bonding interactions with the active site residues Tyr55, His117, and His222. The important role of the nonconserved residues Leu54, His222, Leu306, and Leu308 in inhibitor binding and selectivity was determined by site-directed mutagenesis.
- Medicinal Chemistry and Drug Action, Monash Institute of Pharmaceutical Sciences, Parkville, Victoria, Australia.
Organizational Affiliation: