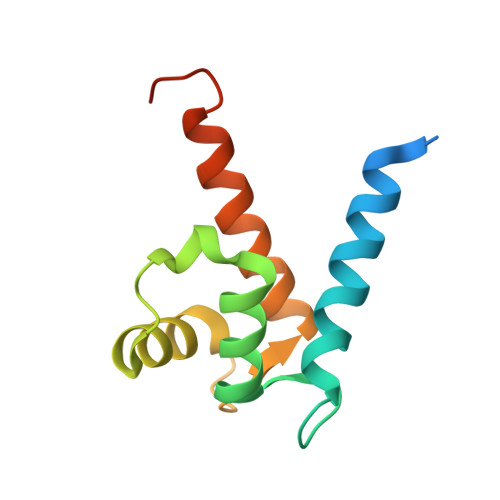
Crystal structure of the Ca(2+)-form and Ca(2+)-binding kinetics of metastasis-associated protein, S100A4
Gingras, A.R., Basran, J., Prescott, A., Kriajevska, M., Bagshaw, C.R., Barsukov, I.L.(2008) FEBS Lett 582: 1651-1656
- PubMed: 18435928 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2008.04.017
- Primary Citation Related Structures:
3C1V - PubMed Abstract:
S100A4 takes part in control of tumour cell migration and contributes to metastatic spread in in vivo models. In the active dimeric Ca(2+)-bound state it interacts with multiple intracellular targets. Conversely, oligomeric forms of S100A4 are linked with the extracellular function of this protein. We report the 1.5A X-ray crystal structure of Ca(2+)-bound S100A4 and use it to identify the residues involved in target recognition and to derive a model of the oligomeric state. We applied stopped-flow analysis of tyrosine fluorescence to derive kinetics of S100A4 activation by Ca(2+) (k(on)=3.5 microM(-1)s(-1), k(off)=20s(-1)).
- Department of Biochemistry, University of Leicester, Henry Wellcome Building, Leicester LE1 9HN, UK.
Organizational Affiliation: