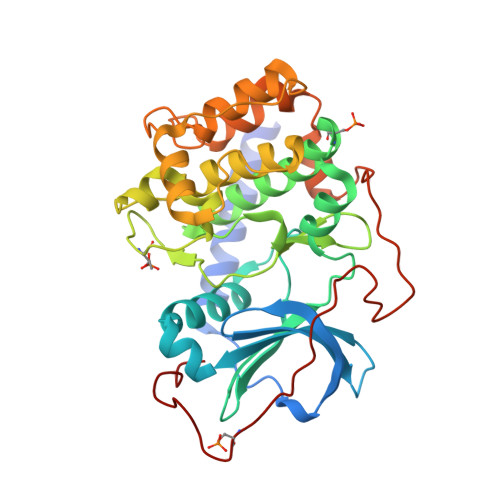
Structural analysis of ARC-type inhibitor (ARC-1034) binding to protein kinase A catalytic subunit and rational design of bisubstrate analogue inhibitors of basophilic protein kinases.
Lavogina, D., Lust, M., Viil, I., Konig, N., Raidaru, G., Rogozina, J., Enkvist, E., Uri, A., Bossemeyer, D.(2009) J Med Chem 52: 308-321
- PubMed: 19143565 Search on PubMed
- DOI: https://doi.org/10.1021/jm800797n
- Primary Citation Related Structures:
3BWJ - PubMed Abstract:
The crystal structure of a complex of the catalytic subunit (type alpha) of cAMP-dependent protein kinase (PKA C alpha) with ARC-type inhibitor (ARC-1034), the presumed lead scaffold of previously reported adenosine-oligo-arginine conjugate-based (ARC-type) inhibitors, was solved. Structural elements important for interaction with the kinase were established with specifically modified derivatives of the lead compound. On the basis of this knowledge, a new generation of inhibitors, conjugates of adenosine-4'-dehydroxymethyl-4'-carboxylic acid moiety and oligo(D-arginine), was developed with inhibitory constants well into the subnanomolar range. The structural determinants of selectivity of the new compounds were established in assays with ROCK-II and PKBgamma.
- Institute of Chemistry, 2 Jakobi Street, 51014 Tartu, Estonia.
Organizational Affiliation: