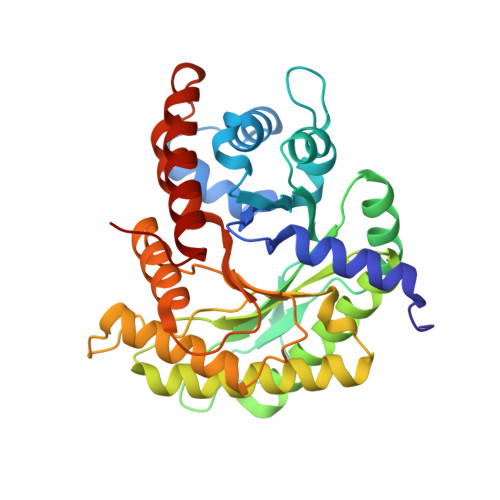
Structure of a rabbit muscle fructose-1,6-bisphosphate aldolase A dimer variant.
Sherawat, M., Tolan, D.R., Allen, K.N.(2008) Acta Crystallogr D Biol Crystallogr 64: 543-550
- PubMed: 18453690 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444908004976
- Primary Citation Related Structures:
3BV4 - PubMed Abstract:
Fructose-1,6-bisphosphate aldolase (aldolase) is an essential enzyme in glycolysis and gluconeogenesis. In addition to this primary function, aldolase is also known to bind to a variety of other proteins, a property that may allow it to perform 'moonlighting' roles in the cell. Although monomeric and dimeric aldolases possess full catalytic activity, the enzyme occurs as an unusually stable tetramer, suggesting a possible link between the oligomeric state and these noncatalytic cellular roles. Here, the first high-resolution X-ray crystal structure of rabbit muscle D128V aldolase, a dimeric form of aldolase mimicking the clinically important D128G mutation in humans associated with hemolytic anemia, is presented. The structure of the dimer was determined to 1.7 angstroms resolution with the product DHAP bound in the active site. The turnover of substrate to produce the product ligand demonstrates the retention of catalytic activity by the dimeric aldolase. The D128V mutation causes aldolase to lose intermolecular contacts with the neighboring subunit at one of the two interfaces of the tetramer. The tertiary structure of the dimer does not significantly differ from the structure of half of the tetramer. Analytical ultracentrifugation confirms the occurrence of the enzyme as a dimer in solution. The highly stable structure of aldolase with an independent active site is consistent with a model in which aldolase has evolved as a multimeric scaffold to perform other noncatalytic functions.
- Department of Physiology and Biophysics, Boston University School of Medicine, 715 Albany Street, Boston, MA 02118-2394, USA.
Organizational Affiliation: