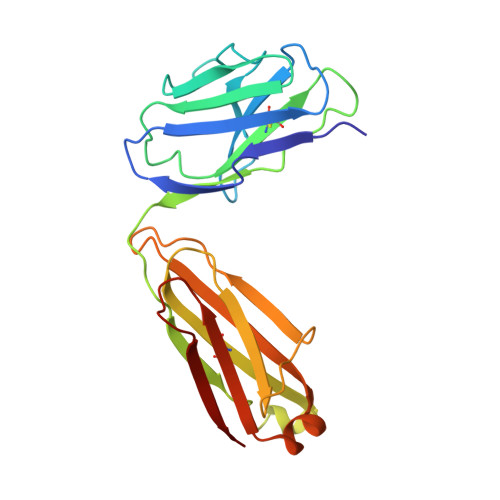
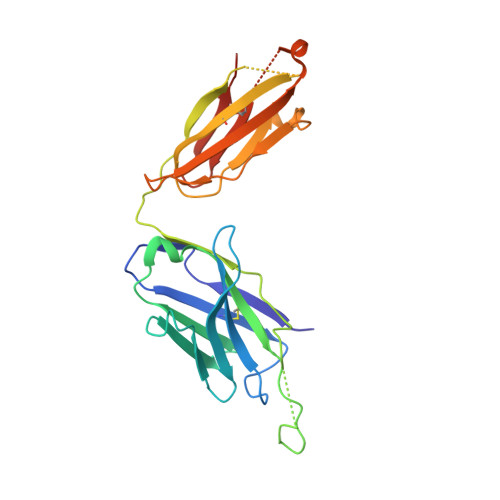
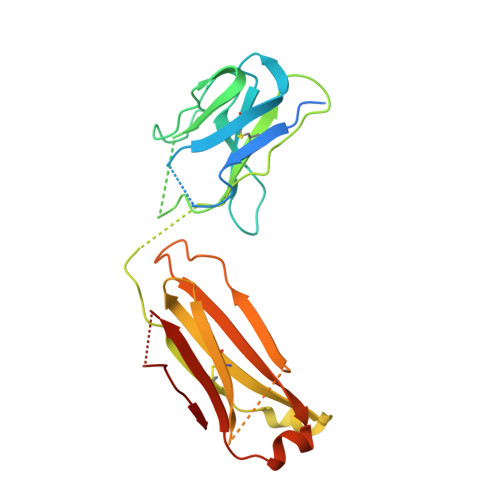
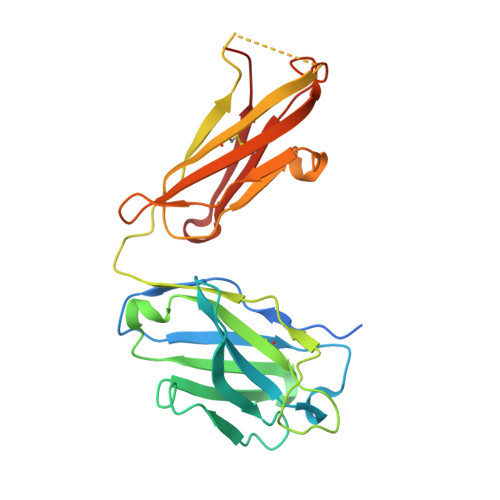
Crystal structure of the complex between the F(ab)' fragment of the cross-neutralizing anti-HIV-1 antibody 2F5 and the F(ab) fragment of its anti-idiotypic antibody 3H6.
Bryson, S., Julien, J.P., Isenman, D.E., Kunert, R., Katinger, H., Pai, E.F.(2008) J Mol Biology 382: 910-919
- PubMed: 18692506 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.07.057
- Primary Citation Related Structures:
3BQU - PubMed Abstract:
The monoclonal antibody 2F5 neutralizes a broad range of human immunodeficiency virus-1 isolates via a conserved epitope on the viral glycoprotein gp41. The conformation of the principal epitope is a type I beta-turn centered on gp41 residues (664)DKW(666); in addition, binding studies indicate that residues N- and C-terminal to this core as well as structurally more distant parts of gp41 also contribute to the interaction. Ab2/3H6 is an anti-idiotypic antibody that inhibits the interaction between 2F5 and gp41 and as such, Ab2/3H6 may, in principle, possess a paratope that mimics the gp41 epitope. To establish the potential of Ab2/3H6 to serve as a guide for the design of vaccine components against human immunodeficiency virus, we investigated the crystal structure of the heterodimeric complex of Ab2/3H6 F(ab) and 2F5 F(ab)'. Ab2/3H6 F(ab) binds to 2F5 F(ab)' via a helix-like protrusion formed by residues (58(H))RYSPSLNTRL(67(H)) of the 2F5 F(ab)' variable domain and proximal to but not overlapping with the gp41 (664)DKW(666) epitope-binding pocket. This defines Ab2/3H6 as an anti-idiotypic antibody of the Ab2gamma class, i.e., an antigen-inhibitable idiotype that does not carry the internal image of the linear primary gp41 (662)ELDKWAS(668) epitope.
- Department of Biochemistry, University of Toronto, 1 King's College Circle, Toronto, Ontario, Canada M5S 1A8.
Organizational Affiliation: