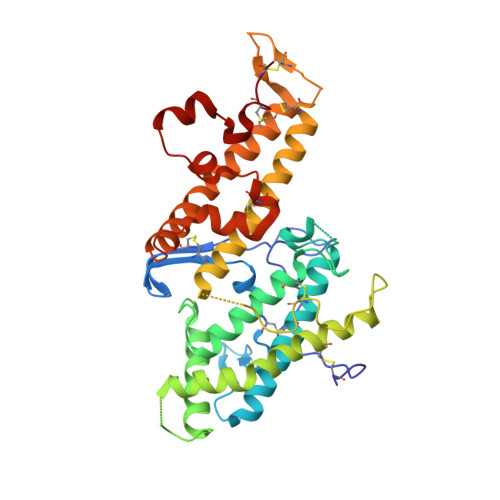
The structure of a chondroitin sulfate-binding domain important in placental malaria.
Higgins, M.K.(2008) J Biological Chem 283: 21842-21846
- PubMed: 18550531 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.C800086200
- Primary Citation Related Structures:
3BQI, 3BQK, 3BQL - PubMed Abstract:
Adhesive PfEMP1 proteins are displayed on the surface of malaria-infected red blood cells. They play a critical role in the disease, tethering infected cells away from destruction by the spleen and causing many severe symptoms. A molecular understanding of how these domains maintain their binding properties while evading immune detection will be important in developing therapeutics. In malaria of pregnancy, domains from the var2csa-encoded PfEMP1 protein interact with chondroitin sulfate on the placenta surface. This causes accumulation of infected red blood cells, leading to placental inflammation and block of blood flow to the developing fetus. This is associated with maternal anemia, low birth weight, and premature delivery and can lead to the death of mother and child. Here I present the structure of the chondroitin sulfate-binding DBL3X domain from a var2csa-encoded PfEMP1 protein. The domain adopts a fold similar to malarial invasion proteins, with extensive loop insertions. One loop is flexible in the unliganded structure but observed in the presence of sulfate or disaccharide, where it completes a sulfate-binding site. This loop, and others surrounding this putative carbohydrate-binding site, are flexible and polymorphic, perhaps protecting the binding site from immune detection. This suggests a model for how the domain maintains ligand binding while evading the immune response and will guide future drug and vaccine development.
- Department of Biochemistry, University of Cambridge, 80, Tennis Court Road, Cambridge CB2 1GA, United Kingdom. mkh20@cam.ac.uk
Organizational Affiliation: