Crystal Structure of HutP complexed with the 55-mer RNA
Kumarevel, T., Balasundaresan, D., Jeyakanthan, J., Shinkai, A., Yokoyama, S., Kumar, P.K.R.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Hut operon positive regulatory protein | B [auth A], C [auth B], D [auth C] | 147 | Bacillus subtilis | Mutation(s): 1 Gene Names: hutP | 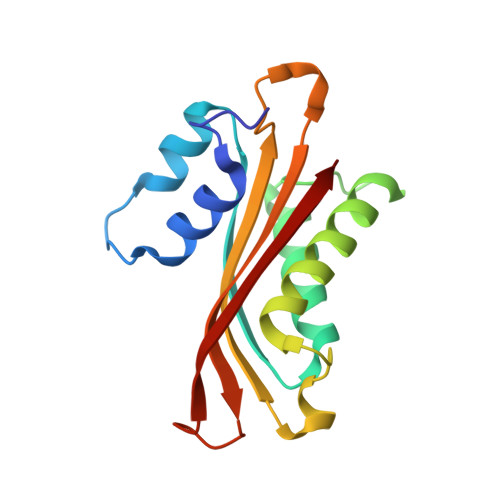 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P10943 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 1 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| 5'-R(*UP*UP*UP*AP*GP*UP*UP*UP*UP*UP*AP*GP*UP*UP*UP*UP*UP*AP*GP*UP*UP*U)-3' | A [auth D] | 22 | N/A | 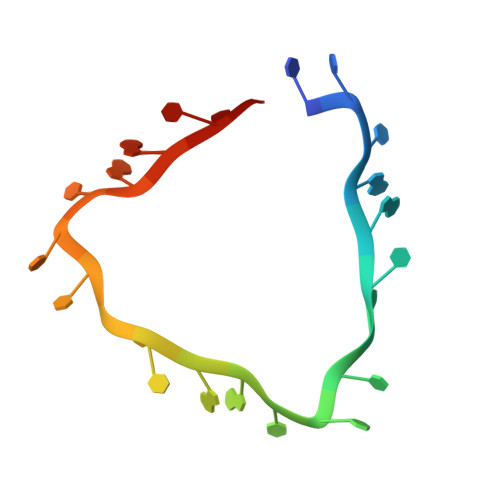 |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| HIS Download:Ideal Coordinates CCD File | F [auth A], G [auth A], H [auth A] | HISTIDINE C6 H10 N3 O2 HNDVDQJCIGZPNO-YFKPBYRVSA-O |  | ||
| MG Download:Ideal Coordinates CCD File | E [auth A], I [auth B], J [auth C] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 99.635 | α = 90 |
| b = 76.469 | β = 109.06 |
| c = 62.79 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| HKL-2000 | data collection |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| MOLREP | phasing |