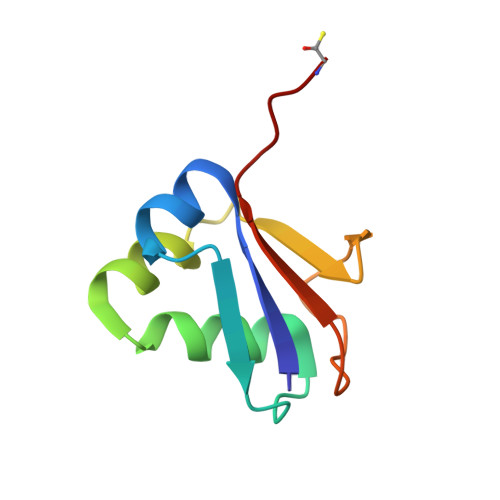
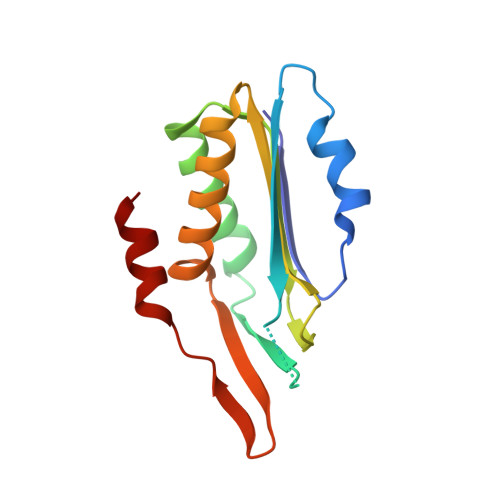
Crystal structure of a molybdopterin synthase-precursor Z complex: insight into its sulfur transfer mechanism and its role in molybdenum cofactor deficiency.
Daniels, J.N., Wuebbens, M.M., Rajagopalan, K.V., Schindelin, H.(2008) Biochemistry 47: 615-626
- PubMed: 18092812 Search on PubMed
- DOI: https://doi.org/10.1021/bi701734g
- Primary Citation Related Structures:
2Q5W, 2QIE, 3BII - PubMed Abstract:
In almost all biological life forms, molybdenum and tungsten are coordinated by molybdopterin (MPT), a tricyclic pyranopterin containing a cis-dithiolene group. Together, the metal and the pterin moiety form the redox reactive molybdenum cofactor (Moco). Mutations in patients with deficiencies in Moco biosynthesis usually occur in the enzymes catalyzing the first and second steps of biosynthesis, leading to the formation of precursor Z and MPT, respectively. The second step is catalyzed by the heterotetrameric MPT synthase protein consisting of two large (MoaE) and two small (MoaD) subunits with the MoaD subunits located at opposite ends of a central MoaE dimer. Previous studies have determined that the conversion of the sulfur- and metal-free precursor Z to MPT by MPT synthase involves the transfer of sulfur atoms from a C-terminal MoaD thiocarboxylate to the C-1' and C-2' positions of precursor Z. Here, we present the crystal structures of non-thiocarboxylated MPT synthase from Staphylococcus aureus in its apo form and in complex with precursor Z. A comparison of the two structures reveals conformational changes in a loop that participates in interactions with precursor Z. In the complex, precursor Z is bound by strictly conserved residues in a pocket at the MoaE dimer interface in close proximity of the C-terminal glycine of MoaD. Biochemical evidence indicates that the first dithiolene sulfur is added at the C-2' position.
- Department of Biochemistry and Cell Biology, Stony Brook University, Stony Brook, New York 11794-5215, USA.
Organizational Affiliation: