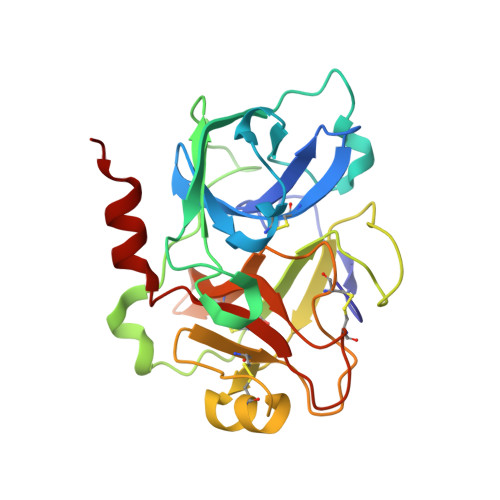
Clavatadine A, a natural product with selective recognition and irreversible inhibition of factor XIa.
Buchanan, M.S., Carroll, A.R., Wessling, D., Jobling, M., Avery, V.M., Davis, R.A., Feng, Y., Xue, Y., Oster, L., Fex, T., Deinum, J., Hooper, J.N., Quinn, R.J.(2008) J Med Chem 51: 3583-3587
- PubMed: 18510371 Search on PubMed
- DOI: https://doi.org/10.1021/jm800314b
- Primary Citation Related Structures:
3BG8 - PubMed Abstract:
Bioassay-guided fractionation of a CH2Cl2/MeOH extract of the sponge Suberea clavata using the serine protease factor XIa to detect antithrombotic activity led to the isolation of the new marine natural products, clavatadines A and B. Clavatadines A and B inhibited factor XIa with IC50's of 1.3 and 27 microM, respectively. A crystal structure of protein-inhibitor (clavatadine A) complex was obtained and revealed interesting selective binding and irreversible inhibition of factor XIa. The cocrystal structure provides guidance for the design and synthesis of future factor XIa inhibitors as antithrombotic agents.
- Eskitis Institute, Griffith University, Brisbane, Queensland 4111, Australia.
Organizational Affiliation: