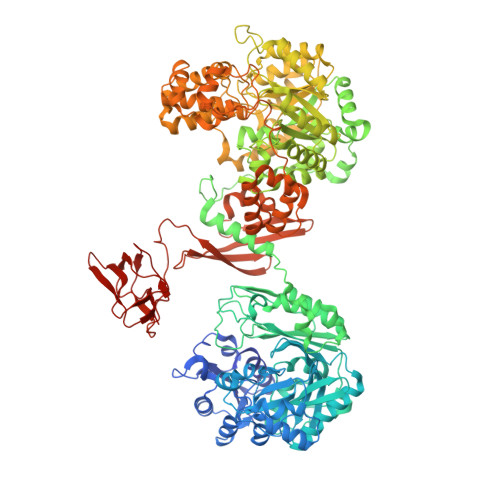
Crystal structures of human and Staphylococcus aureus pyruvate carboxylase and molecular insights into the carboxyltransfer reaction.
Xiang, S., Tong, L.(2008) Nat Struct Mol Biol 15: 295-302
- PubMed: 18297087 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.1393
- Primary Citation Related Structures:
3BG3, 3BG5, 3BG9 - PubMed Abstract:
Pyruvate carboxylase (PC) catalyzes the biotin-dependent production of oxaloacetate and has important roles in gluconeogenesis, lipogenesis, insulin secretion and other cellular processes. PC contains the biotin carboxylase (BC), carboxyltransferase (CT) and biotin-carboxyl carrier protein (BCCP) domains. We report here the crystal structures at 2.8-A resolution of full-length PC from Staphylococcus aureus and the C-terminal region (missing only the BC domain) of human PC. A conserved tetrameric association is observed for both enzymes, and our structural and mutagenesis studies reveal a previously uncharacterized domain, the PC tetramerization (PT) domain, which is important for oligomerization. A BCCP domain is located in the active site of the CT domain, providing the first molecular insights into how biotin participates in the carboxyltransfer reaction. There are dramatic differences in domain positions in the monomer and the organization of the tetramer between these enzymes and the PC from Rhizobium etli.
- Department of Biological Sciences, 212 Amsterdam Avenue, Columbia University, New York, New York 10027, USA.
Organizational Affiliation: