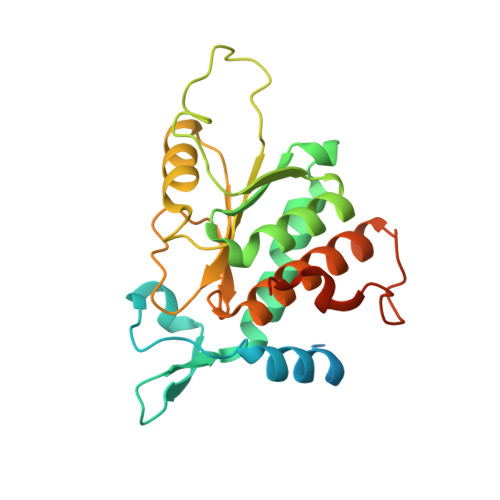
Crystal structure of the human laminin receptor precursor.
Jamieson, K.V., Wu, J., Hubbard, S.R., Meruelo, D.(2008) J Biological Chem 283: 3002-3005
- PubMed: 18063583 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.C700206200
- Primary Citation Related Structures:
3BCH - PubMed Abstract:
The human laminin receptor (LamR) interacts with many ligands, including laminin, prions, Sindbis virus, and the polyphenol (-)-epigallocatechin-3-gallate (EGCG), and has been implicated in a number of diseases. LamR is overexpressed on tumor cells, and targeting LamR elicits anti-cancer effects. Here, we report the crystal structure of human LamR, which provides insights into its function and should facilitate the design of novel therapeutics targeting LamR.
- Gene Therapy Center, Cancer Institute and Department of Pathology, New York University School of Medicine, New York, New York 10016.
Organizational Affiliation: