Novel thiol-based TACE inhibitors. Part 2: Rational design, synthesis, and SAR of thiol-containing aryl sulfones.
Bandarage, U.K., Wang, T., Come, J.H., Perola, E., Wei, Y., Rao, B.G.(2008) Bioorg Med Chem Lett 18: 44-48
- PubMed: 18054488 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2007.11.014
- Primary Citation Related Structures:
3B92 - PubMed Abstract:
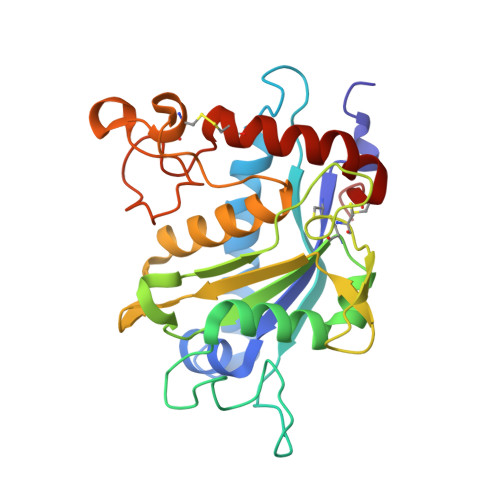
A series of potent thiol-containing aryl sulfone TACE inhibitors were designed and synthesized. The SAR and MMP selectivity of the series were investigated. In particular, compound 8b showed excellent in vitro potency against the isolated enzyme and good selectivity over MMP-2, -7, -8, -9, and -13. The X-ray structure of 8b in complex with TACE was also obtained.
- Vertex Pharmaceuticals Incorporated, 130 Waverly Street, Cambridge, MA 02139, USA. upul_bandarage@vrtx.com
Organizational Affiliation: