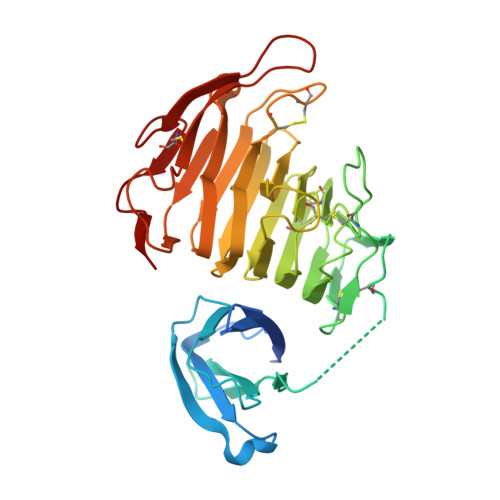
The Crystal Structure of Pectate Lyase PelI from Soft Rot Pathogen Erwinia chrysanthemi in Complex with Its Substrate.
Creze, C., Castang, S., Derivery, E., Haser, R., Hugouvieux-Cotte-Pattat, N., Shevchik, V.E., Gouet, P.(2008) J Biological Chem 283: 18260-18268
- PubMed: 18430740 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M709931200
- Primary Citation Related Structures:
3B4N, 3B8Y, 3B90 - PubMed Abstract:
The crystallographic structure of the family 3 polysaccharide lyase (PL-3) PelI from Erwinia chrysanthemi has been solved to 1.45 A resolution. It consists of an N-terminal domain harboring a fibronectin type III fold linked to a catalytic domain displaying a parallel beta-helix topology. The N-terminal domain is located away from the active site and is not involved in the catalytic process. After secretion in planta, the two domains are separated by E. chrysanthemi proteases. This event turns on the hypersensitive response of the host. The structure of the single catalytic domain determined to 2.1 A resolution shows that the domain separation unveils a "Velcro"-like motif of asparagines, which might be recognized by a plant receptor. The structure of PelI in complex with its substrate, a tetragalacturonate, has been solved to 2.3 A resolution. The sugar binds from subsites -2 to +2 in one monomer of the asymmetric unit, although it lies on subsites -1 to +3 in the other. These two "Michaelis complexes" have never been observed simultaneously before and are consistent with the dual mode of bond cleavage in this substrate. The bound sugar adopts a mixed 2(1) and 3(1) helical conformation similar to that reported in inactive mutants from families PL-1 and PL-10. However, our study suggests that the catalytic base in PelI is not a conventional arginine but a lysine as proposed in family PL-9.
- Laboratoire de BioCristallographie, Institut de Biologie et Chimie des Protéines, CNRS et Université de Lyon, UMR 5086, IFR 128 BioSciences Gerland-Lyon Sud, Lyon, France.
Organizational Affiliation: