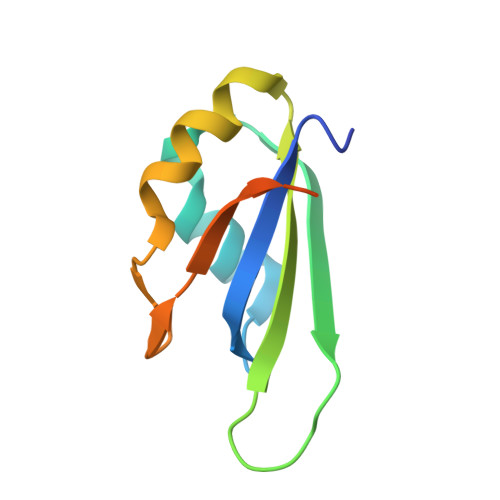
Crystal structure and possible dimerization of the single RRM of human PABPN1
Ge, H., Zhou, D., Tong, S., Gao, Y., Teng, M., Niu, L.(2008) Proteins 71: 1539-1545
- PubMed: 18275081 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21973
- Primary Citation Related Structures:
3B4D, 3B4M - Hefei National Laboratory for Physical Sciences at Microscale, School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230026, China.
Organizational Affiliation: