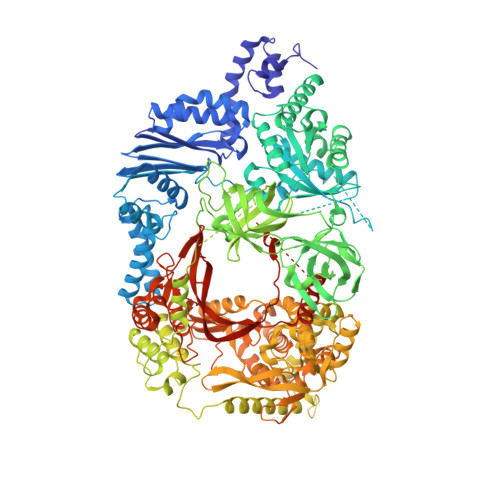
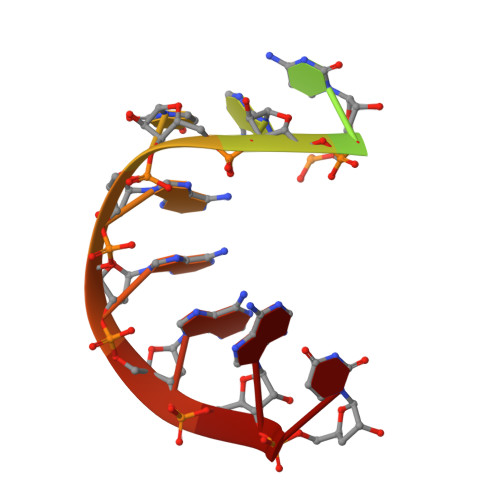
Molecular basis for RNA polymerization by Q beta replicase
Takeshita, D., Tomita, K.(2012) Nat Struct Mol Biol 19: 229-237
- PubMed: 22245970 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2204
- Primary Citation Related Structures:
3AVT, 3AVU, 3AVV, 3AVW, 3AVX, 3AVY - PubMed Abstract:
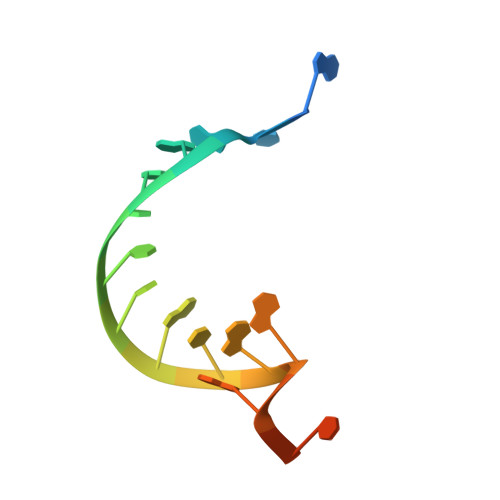
Core Qβ replicase comprises the Qβ virus-encoded RNA-dependent RNA polymerase (β-subunit) and the host Escherichia coli translational elongation factors EF-Tu and EF-Ts. The functions of the host proteins in the viral replicase are not clear. Structural analyses of RNA polymerization by core Qβ replicase reveal that at the initiation stage, the 3'-adenine of the template RNA provides a stable platform for de novo initiation. EF-Tu in Qβ replicase forms a template exit channel with the β-subunit. At the elongation stages, the C-terminal region of the β-subunit, assisted by EF-Tu, splits the temporarily double-stranded RNA between the template and nascent RNAs before translocation of the single-stranded template RNA into the exit channel. Therefore, EF-Tu in Qβ replicase modulates RNA elongation processes in a distinct manner from its established function in protein synthesis.
- Biomedical Research Institute, National Institute of Advanced Industrial Science and Technology, Ibaraki, Japan.
Organizational Affiliation: