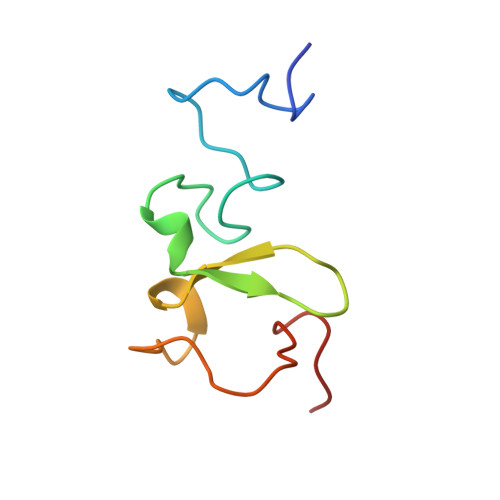
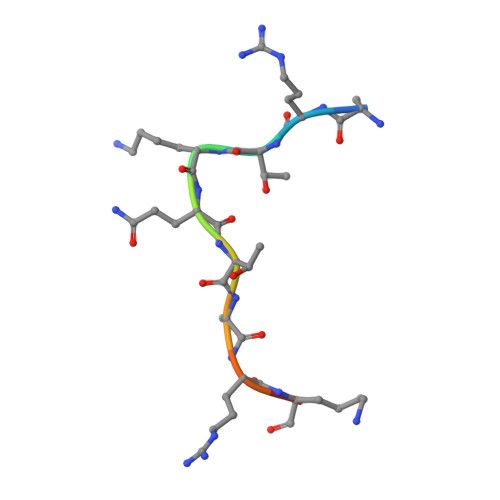
Recognition of modification status on a histone H3 tail by linked histone reader modules of the epigenetic regulator UHRF1
Arita, K., Isogai, S., Oda, T., Unoki, M., Sugita, K., Sekiyama, N., Kuwata, K., Hamamoto, R., Tochio, H., Sato, M., Ariyoshi, M., Shirakawa, M.(2012) Proc Natl Acad Sci U S A 109: 12950-12955
- PubMed: 22837395 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1203701109
- Primary Citation Related Structures:
3ASK, 3ASL - PubMed Abstract:
Multiple covalent modifications on a histone tail are often recognized by linked histone reader modules. UHRF1 [ubiquitin-like, containing plant homeodomain (PHD) and really interesting new gene (RING) finger domains 1], an essential factor for maintenance of DNA methylation, contains linked two-histone reader modules, a tandem Tudor domain and a PHD finger, tethered by a 17-aa linker, and has been implicated to link histone modifications and DNA methylation. Here, we present the crystal structure of the linked histone reader modules of UHRF1 in complex with the amino-terminal tail of histone H3. Our structural and biochemical data provide the basis for combinatorial readout of unmodified Arg-2 (H3-R2) and methylated Lys-9 (H3-K9) by the tandem tudor domain and the PHD finger. The structure reveals that the intermodule linker plays an essential role in the formation of a histone H3-binding hole between the reader modules by making extended contacts with the tandem tudor domain. The histone H3 tail fits into the hole by adopting a compact fold harboring a central helix, which allows both of the reader modules to simultaneously recognize the modification states at H3-R2 and H3-K9. Our data also suggest that phosphorylation of a linker residue can modulate the relative position of the reader modules, thereby altering the histone H3-binding mode. This finding implies that the linker region plays a role as a functional switch of UHRF1 involved in multiple regulatory pathways such as maintenance of DNA methylation and transcriptional repression.
- Department of Molecular Engineering, Graduate School of Engineering, Kyoto University, Kyoto 615-8510, Japan.
Organizational Affiliation: